Transforming Histology into Virtual Multiplex Immunofluorescence to Decode Prognostic Spatial Immunity in Hepatocellular Carcinoma
Transforming Histology into Virtual Multiplex Immunofluorescence to Decode Prognostic Spatial Immunity in Hepatocellular Carcinoma
Cai, L.; Jiang, S.; Liang, J.; Liu, F.; Zhang, B.; Reitsam, N. G.; Zeng, Q.; Ma, Y.; Li, Z.; Feng, S.; Hu, M.; Zhang, X.; Zhang, J.; Kather, J. N.; Zhang, Y.; Liang, W.
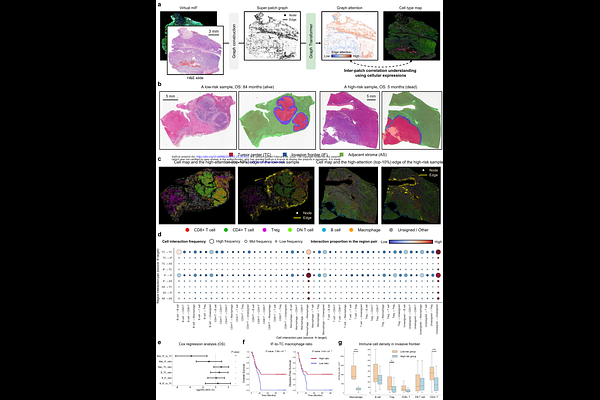
AbstractThe spatial organization of the tumor immune microenvironment (TIME) drives hepatocellular carcinoma (HCC) prognosis but remains unquantifiable on routine H&E slides. Here, we present HCCExplorer, a deep learning framework that translates H&E into virtual multiplex immunofluorescence (mIF) and uses multi-modal graph learning to decode spatial survival signals. Trained on 30 H&E-mIF slide pairs, HCCExplorer evaluated a 1,813-slide multi-center cohort. It achieved superior overall survival stratification over clinical indices and state-of-the-art pathology and protein foundation models, yielding a concordance index of 0.71 and a Hazard Ratio (HR) of 15.46 (P<0.001), maintaining stability across three external cohorts. Beyond risk stratification, interpretation of model features identified M1 macrophage infiltration as a protective determinant (HR=0.40, P<0.05). Furthermore, it uncovered a protective Containment Niche at the invasion frontier (HR=0.02, P<0.01), featuring macrophages co-localizing with Foxp3+ Tregs and CD4+ T cells. Ultimately, HCCExplorer provides actionable, spatially-resolved biomarkers from conventional histology for precision HCC management.