BayesR3AD: Joint analysis of additive and dominance in Bayesian mixture models
BayesR3AD: Joint analysis of additive and dominance in Bayesian mixture models
Yuan, H.; Breen, E. J.; MacLeod, I. M.; Khansefid, M.; Xiang, R.; Goddard, M. E.
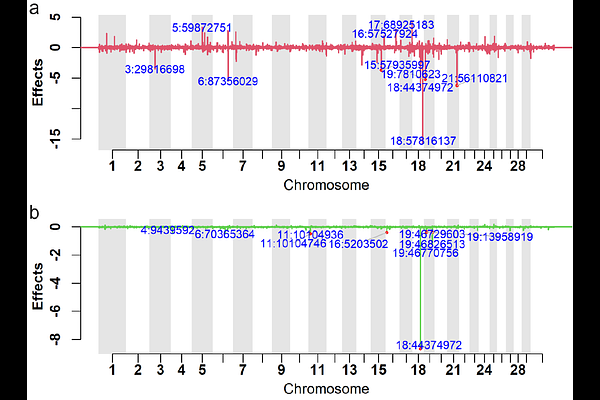
AbstractBackground: Genomic prediction in livestock is predominantly based on additive models, even though dominance and other non-additive effects can contribute appreciably to phenotypic variance for fitness and fertility traits. Bayesian mixture models, such as Bayes R, have proven effective for modelling sparse, heterogeneous additive SNP effects, but most implementations do not explicitly accommodate dominance. In this study, we extended BayesR3 to jointly model additive and dominance marker effects within a unified Bayesian mixture framework, denoted BayesR3AD, and used this method to estimate additive and dominance effects for fertility and cow survival (longevity) in Holstein cattle. Results: Using real Holstein genotypes (227,942 animals, 74,626 SNPs), we simulated phenotypes with additive and dominance effects. When dominance was present in the simulated data, BayesR3AD improved prediction accuracy of genetic values by +0.1011 (0.6144 vs 0.5133; {approx}19.7% relative) compared with the additive-only BayesR3 model and recovered additive and dominance variance components without bias. Under purely additive simulations, dominance mixture components were effectively empty, confirming that the extended model shrinks unnecessary dominance effects toward zero. In real fertility data, including calving interval (63,378 records) and survival (68,514 records), BayesR3AD estimated small dominance variance ({approx}1-3% of total genetic variance). The model highlighted a very large additive loci at 57.82 Mb on BTA18 for both calving interval and survival. concordant with previous GWAS studies of Holstein fertility. Additionally, a large dominance effect was found at 44.37 Mb on BTA18 for calving interval implicating a heterozygote advantage that increases fertility. Conclusions: BayesR3AD provides a practical extension of BayesR3 that captures both additive and dominance contributions to genomic prediction. The method is robust, reverting effectively to the additive model when dominance is absent, while delivering accurate variance decomposition, and potential gains in prediction accuracy when dominance is present. Application to Holstein fertility traits demonstrates that dominance can be detected and quantified without compromising additive inference, supporting improved prediction of total genetic merit. While validated in cattle, BayesR3AD can be directly applied to other species to better model and predict traits related to fitness.