Genomic diversity of British native oaks: species differentiation, hybridisation and triploidy
Genomic diversity of British native oaks: species differentiation, hybridisation and triploidy
Gathercole, L. A.; Carleial, R.; Brown, N.; Denman, S.; Wu, E.; Nichols, R. A.; Buggs, R. J. A.
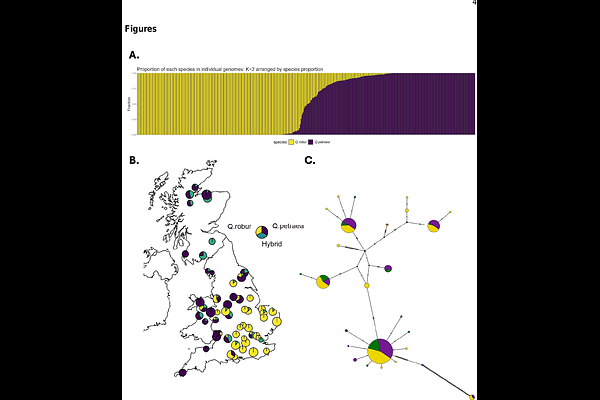
AbstractThe two native British oak species Quercus robur (pedunculate oak) and Q. petraea (sessile oak) are widespread and of ecological, cultural and economic value. Here we sequence whole genomic DNA of 418 individuals from 60 natural oak populations, most of which were previously monitored in a Forest Condition Survey that ran from 1987 to 2007 across Great Britain. Variant calling identifies 146,995,863 single nucleotide polymorphisms (SNPs) and 28,275,691 indels. Analysing population genetic structure at 7,112,139 biallelic SNPs, we find two groups, indicating that Q. robur is more common in the south and east of Britain, and Q. petraea in the north and west. The species distributions show niche differentiation in that Q. robur prefers warmer and more thermally variable environments with alkaline soils, while Q. petraea prefers rainy environments with more topographically complex terrain. We show extensive hybridisation and back-crossing between the two species. This is biased in favour of introgression from Q. robur into Q. petraea. By analysis of allele balance, we find five triploid trees among our samples, two belonging to each species and one being hybridised. We find little population genetic structure within species. Despite genome-wide introgression, we find islands of genomic differentiation between species, particularly in chromosome two. These differentiated regions contain 8,230 SNPs of which 2,005 are within gene annotations. We also show widespread chloroplast haplotype sharing between species. Between 1990 and 2019, mean stem growth was higher in Q. robur than in Q. petraea and hybrids, due to environmental factors. Triploids grew significantly faster than diploids even when environmental factors were accounted for.