FLIP2: Expanding Protein Fitness Landscape Benchmarks for Real-World Machine Learning Applications
FLIP2: Expanding Protein Fitness Landscape Benchmarks for Real-World Machine Learning Applications
Didi, K.; Alamdari, S.; Lu, A. X.; Wittmann, B.; Johnston, K. E.; Amini, A. P.; Madani, A. K.; Czeneszew, M.; Dallago, C.; Yang, K. K.
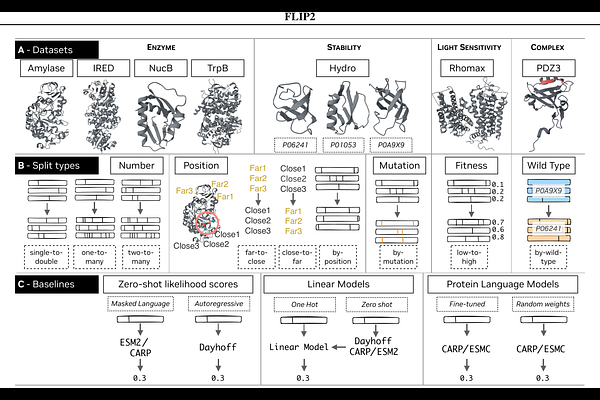
AbstractMachine learning methods that predict protein fitness from sequence remain sensitive to changes in data distributions, limiting generalization across common conditions encountered in protein engineering. Practically, protein engineers are thus left wondering about the effective utility of ML tools. The FLIP benchmark established protocols for testing generalization under some domain shifts, but it was limited to measurements of thermostability, binding, and viral capsid viability. We introduce FLIP2, a protein fitness benchmark spanning seven new datasets, including enzymes, protein-protein interactions, and light-sensitive proteins, as well as splits that measure generalization relevant to real-world protein engineering campaigns. Evaluating a suite of benchmark models across these datasets and suites reveals that simpler models often matched or outperformed fine-tuned protein language models on \ourset, challenging the utility of existing transfer learning techniques. Provenance for all datasets has been recorded and we redistribute all data CC-BY 4.0 to facilitate continued progress.