BertMS-enabled molecular networking for unknown compounds dereplication
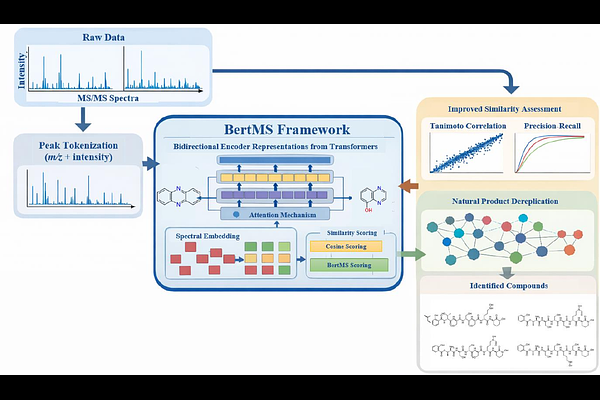
BertMS-enabled molecular networking for unknown compounds dereplication
Luning, Z.; Shuang, W.; Jixing, P.; Xiaofei, H.; Wenxue, W.; Dehai, L.
AbstractSpectral similarity is widely used as a proxy for structural similarity in tandem mass spectrometry (MS/MS) analyses, including library matching and molecular networking. However, the relationship between spectral similarity scores and true structural similarity remains imperfect, limiting compound identification in metabolomics studies. Here, we present BertMS, a spectral similarity framework based on bidirectional encoder representations from transformers (BERT), which learns contextualized representations of fragment ions from large scale MS/MS data. Using datasets from MoNA and GNPS comprising over 100,000 unique molecules, we systematically evaluate BertMS against existing methods, including cosine similarity and Spec2Vec. BertMS shows improved performance across multiple evaluation metrics, with average gains of approximately 15-25% depending on the task. Notably, improvements are most evident in molecular similarity assessment. We further demonstrate the applicability of BertMS in molecular networking and dereplication of microbial metabolites, where it enables more consistent identification of structurally related compounds. These results suggest that transformer-based representations provide a useful framework for improving spectral similarity estimation and facilitating metabolite annotation in complex mixtures.