Isoform-resolved spatial transcriptomics on a lab-made high-density array via a single-chip NGS-TGS workflow
Isoform-resolved spatial transcriptomics on a lab-made high-density array via a single-chip NGS-TGS workflow
Yue, Z.; Liu, M.; Liu, Y.; Lu, D.; Zhang, M.; Wang, Y.; Shi, Y.; Miao, Y.; Wang, S.; Jiang, Y.; Wang, Y.; Zhao, J.; Liu, N.; Lv, C.; Zhai, J.; Li, B.
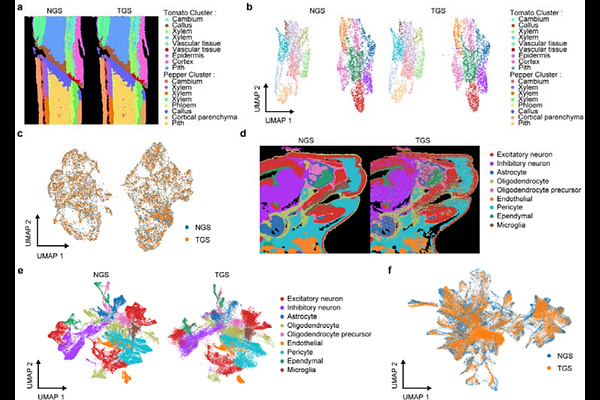
AbstractHigh-resolution spatial transcriptomics is still cost-prohibitive and dominated by short-read sequencing, limiting in situ detection of transcript structures. Here we present a low-cost, lab-made, high-density bead-in-microwell chip assembled by routine centrifuge and decoded by a tri-part combinatorial long-barcode strategy. The design provides 56 million barcode combinations across 4.1 million capture sites, reducing collisions while remaining tolerant to third-generation sequencing raw errors. Using a single-chip, dual-platform workflow, we split full-length cDNA for next-generation and third-generation sequencing within the same spatial coordinates. In an incompatible tomato-pepper graft, we identified interface-associated splicing reprogramming, including thousands of unannotated isoforms, enriched novel CaGRP1 isoforms, and SlPIP2 intron retention, supported by individual long-read sequences. In mouse embryos, we further demonstrate cross-species applicability by resolving unannotated Col1a2 isoforms in oligodendrocyte progenitors and spatially restricted Dalrd3 intron retention. This cost-efficient system broadens access to spatial full-length transcriptomics and may support diverse biological studies.