Comparative cross-species transcriptomic analysis identifies new candidates of Pooideae nitrate response
Comparative cross-species transcriptomic analysis identifies new candidates of Pooideae nitrate response
Gregoire, M.; Pateyron, S.; Brunaud, V.; Tamby, J. P.; Benghelima, L.; Martin, M.-L.; Girin, T.
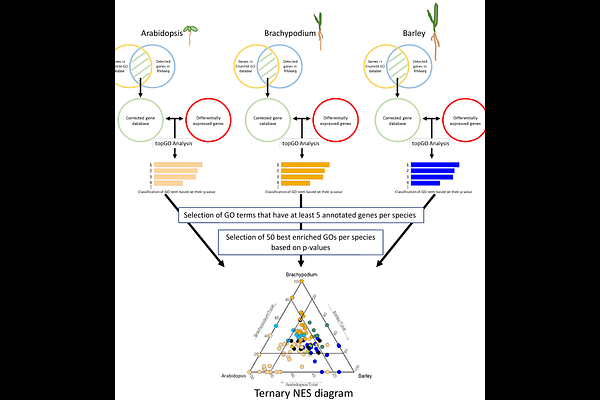
AbstractNitrogen fertilizers are essential for crop productivity but cause environmental harm, necessitating the development of cultivars that thrive under limited nitrogen. This study investigates the transcriptomic response to nitrate in Arabidopsis thaliana (a model dicot), Brachypodium distachyon (a model Pooideae), and Hordeum vulgare (barley, a domesticated Pooideae) to identify conserved and species-specific molecular mechanisms. Using RNA-seq after 1.5 and 3 hours of nitrate treatment, we found that core nitrate-responsive biological processes - such as nitrate transport, assimilation, carbon metabolism, and hormone signaling - are largely conserved across species. However, comparative analysis at gene level based on orthology revealed specificities between the species. For instance, rRNA processing was uniquely stimulated in Arabidopsis, while cysteine biosynthesis from serine and gibberellin biosynthesis were specifically regulated in Brachypodium and barley. Orthologs of key nitrate-responsive genes (e.g., NRT, NLP, TCP20) exhibited variable regulation, reflecting potential adaptations linked to domestication or nutrient acquisition strategies. These findings highlight the importance of integrating model and crop species to uncover targets for improving nitrogen use efficiency in cereals. The study provides a pipeline integrating gene ontology and orthology analyses to compare transcriptomic responses between species.