Decoding epitope immunodominance in HIV Env using cryoEM and machine learning
Decoding epitope immunodominance in HIV Env using cryoEM and machine learning
Schuhmacher, J.; Xiao, S.; Eray, E. R.; Brown, S.; Zambrowski, A.; Jain, A.; Garcia, D. M.; Ozorowski, G.; Zhu, W.; Saam, K.; Caniels, T. G.; Moore, J. P.; Crispin, M.; Sanders, R. W.; Chakraborty, S.; Correia, B. E.; Ward, A. B.; Antanasijevic, A.
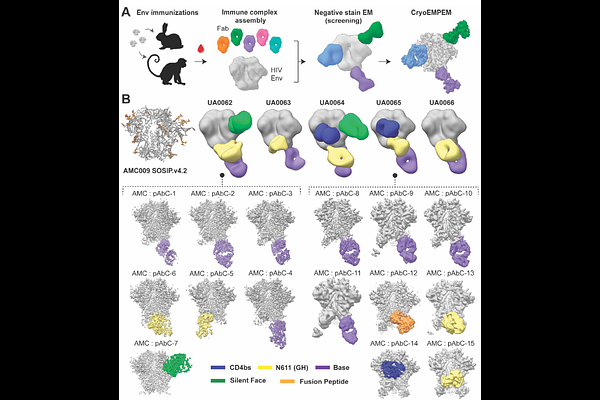
AbstractViral surface glycoproteins, such as the HIV envelope protein (Env), present numerous antibody (Ab) epitopes, yet immune responses consistently focus on only a subset, a phenomenon known as immunodominance. Although structural studies have provided insights into Env antigenicity, our understanding of the molecular features that govern efficient Ab engagement remains incomplete, thereby limiting the predictive and rational design of vaccines. Here, we characterized the structural determinants of epitope immunodominance in HIV Env by integrating high-resolution cryoEM-based polyclonal epitope mapping (cryoEMPEM) across different clades with quantitative analyses of epitope topology, accessibility, and physicochemical properties. More than 70 new structures were resolved to assemble a library of >100 Env-antibody complexes. These data informed the development of a surface-centric, machine-learning model to predict relative Antigen Surface Immunodominance (ASI model). Comparison of ASI-predicted epitope sites with the specificities of Env-induced antibodies showed that the model accurately identifies immunodominant regions and highlights the structural features driving immune bias. Notably, immunogens redesigned based on model predictions successfully redirected Ab responses toward a normally subdominant epitope, demonstrating the potential of strategies coupling targeted assembly of focused structural libraries with machine learning to uncover complex molecular patterns and enable design of more effective vaccine antigens.