Multiomic Spatial Imaging Assay (MSIA) -- A High-plex In Situ Detection Method for mRNAs, Proteins, and Protein-Protein Interactions using Manual And Semi-Automated Workflows
Multiomic Spatial Imaging Assay (MSIA) -- A High-plex In Situ Detection Method for mRNAs, Proteins, and Protein-Protein Interactions using Manual And Semi-Automated Workflows
Zhou, C.; Li, Z.; Zhang, J.; Wang, Y.; Liu, Y.; Williamson, P.; Kim, G.-A.; Deshpande, S. A.; Yuan, M.; Chandrababu, S.; Duan, L.; Chang, C.-W.; Wang, L.-C.; Srinivasan, M.
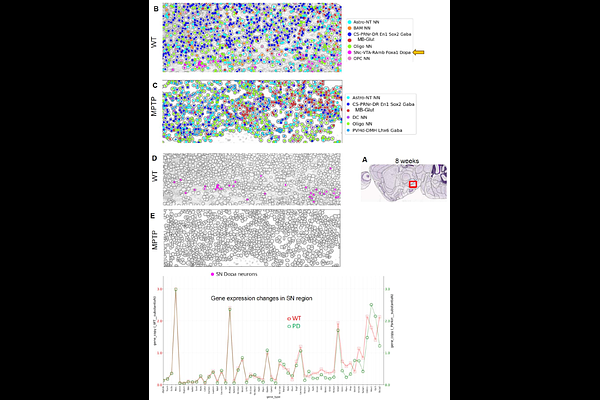
AbstractWe report Multiomics Spatial Image Analysis (MSIA), a suite of technologies that include streamlined manual and semi-automated workflows to image 100s of RNAs from fresh frozen and formalin-fixed paraffin-embedded (FFPE) samples, contextualize RNAs with select protein markers including imaging synaptic protein interactions, a data analysis pipeline that includes corrections for chromatic aberration, algorithms for image stitching, deep learning models for image analysis (cell segmentation and RNA dot detection using hundreds of RNAscope images) from multiple stain-image cycles and proof of concept for a language model trained on extensive Parkinson's Disease (PD) literature. We used the language model to identify potential new biomarkers that were confirmed in MPTP-treated mouse model of PD. Furthermore, by incorporating more stain-image cycles alongside an additional fluorescent channel and advanced error-correcting decoding schemes, the MSIA workflow demonstrates the scalability to potentially quantify over 1,000 genes.