GROQ-seq Enables Cross-site Reproducibility for High-Throughput Measurement of Protein Function
GROQ-seq Enables Cross-site Reproducibility for High-Throughput Measurement of Protein Function
Spinner, A.; Ross, D.; Cortade, D.; Ikonomova, S.; Baranowski, C.; Dhroso, A.; Reider Apel, A.; Sheldon, K.; Duquette, C.; Kelly, P. J.; DeBenedictis, E.; Hudson, C.
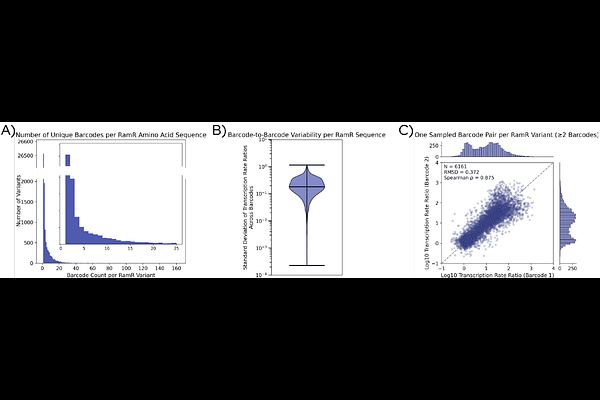
AbstractHigh-throughput functional assays are increasingly used to generate large-scale protein function datasets for protein engineering and machine learning applications. However, the utility of such datasets depends on the reproducibility of the underlying measurements. Here we report reproducible, quantitative measurements of protein sequence-to-function data at scale across two facilities. We analyze GROQ-seq (Growth-based Quantitative Sequencing) measurements of three bacterial transcription factors. Independent barcode measurements of the same sequence produce highly consistent functional estimates, demonstrating strong biological reproducibility (across all transcription factors the mean Root Mean Square Deviation [RMSD] {approx} 0.53 and mean Spearman {approx} 0.63). We also compared experiments performed at two facilities using a shared protocol, but with differing levels of automation and system integration. We observe strong agreement between measurements taken at the two sites (mean RMSD {approx} 0.41 and mean Spearman {approx} 0.730). Orthogonal tests further support this agreement: a classifier trained to distinguish data by site performs near random (AUC = 0.559), and top-ranking variants show strong statistical overlap between experiments. Together, these results demonstrate that GROQ-seq enables reproducible, scalable measurement of protein function suitable for large aggregated datasets.