Beyond Exons: Linking Noncoding Heritability and Polygenicity across Complex Human Traits and Disorders
Beyond Exons: Linking Noncoding Heritability and Polygenicity across Complex Human Traits and Disorders
Fuhrer, J.; Shadrin, A. A.; Hughes, T.; Parker, N.; Hindley, G.; Frei, E.; Nguyen, D.; Smeland, O. B.; Djurovic, S.; Andreassen, O.; Dale, A.; Frei, O.
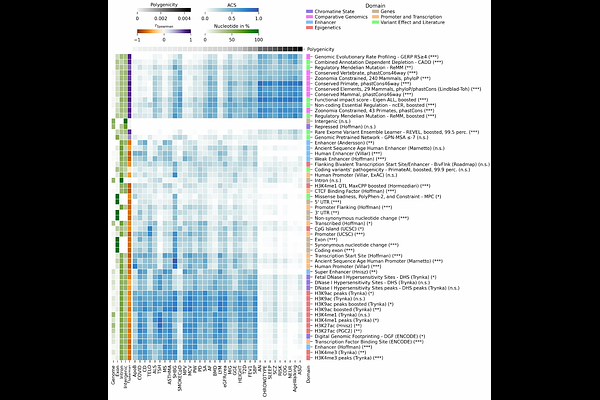
AbstractThe genetic architecture of complex traits spans a continuum of polygenicity, yet it remains unclear how differences in polygenicity relate to the functional localization of SNP heritability across the genome. We use a MiXeR-based framework to partition heritability across exonic, intronic, and intergenic regions for 34 traits and introduce a likelihood-based annotation contribution score that quantifies annotation-specific impact on heritability. Exons explain a minority of heritability, and their contribution decreases with increasing polygenicity, from an average of 22% in less polygenic somatic diseases and biomarkers to 13% in highly polygenic psychiatric and cognitive phenotypes. Intergenic fractions show the opposite trend, whereas intronic fractions remain relatively stable. Analysis of a broader set of functional annotations reveals systematic differences along the polygenicity axis: highly polygenic traits show stronger contributions from comparative genomics and variant-effect scores, whereas less polygenic traits show stronger contributions in promoter, transcription, and chromatin annotations. Together, these results indicate that the functional partitioning of heritability systematically varies with polygenicity, pointing to a shift from gene-proximal regulatory architectures to architectures shaped by numerous dispersed regulatory effects as a key determinant of differences in polygenicity across traits.