An Empirical Bayes approach for the study of phenotypic evolution from high-dimensional data
An Empirical Bayes approach for the study of phenotypic evolution from high-dimensional data
Montoya, P.; Fabre, A.-C.; Goswami, A.; Morlon, H.; Clavel, J.
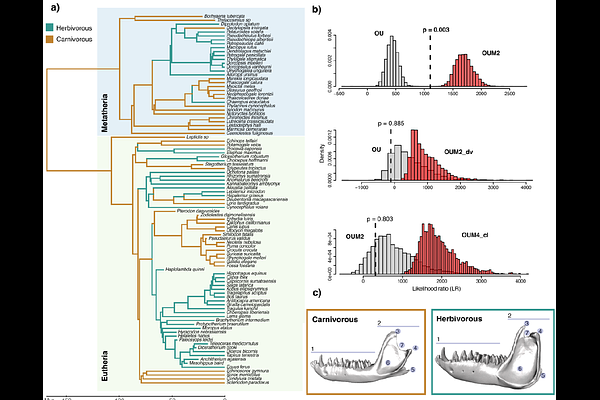
AbstractMultivariate phylogenetic comparative methods for modelling high-dimensional traits such as 3D shapes or gene expression profiles have been recently developed. However, these approaches are impractical and almost impossible to use when the number of traits exceeds a few thousands, as they become computationally prohibitive. We overcome these limitations by proposing a new maximum likelihood approach based on the Empirical Bayes framework. This approach takes into account the information of the complete covariances (among species and traits) to infer parameters and compare models of trait evolution for high-dimensional datasets. Through simulations, we demonstrate that the proposed approach can accurately estimate parameters of various trait evolution models, even when the number of traits is ten times larger than the number of lineages; it requires less memory and is at least 10 times faster than currently available approaches. This fast, efficient framework enabled us to extend the high-dimensional multivariate phylogenetic comparative toolkit by including an Ornstein-Uhlenbeck process with multiple optima to study adaptation to various selective regimes. Applying our approach to the evolution of jaw morphology in relation to dietary adaptation in mammals, we demonstrate morphological convergence in carnivorous and herbivorous lineages. The proposed Empirical Bayes framework, implemented in the R package mvMORPH, enables phylogenetic comparative methods to efficiently handle high-dimensional datasets and complex models of trait evolution.