Specialization of independently acquired flagellar FliC proteins in plant-associated Sphingomonas balances swimming and immunogenicity
Specialization of independently acquired flagellar FliC proteins in plant-associated Sphingomonas balances swimming and immunogenicity
Russ, D.; Saha, C.; Paul, K.; Zheng, Z.; Law, T. F.; Anguita-Maeso, M.; Lundberg, D. S.; Fitzpatrick, C. R.; Dangl, J. L.
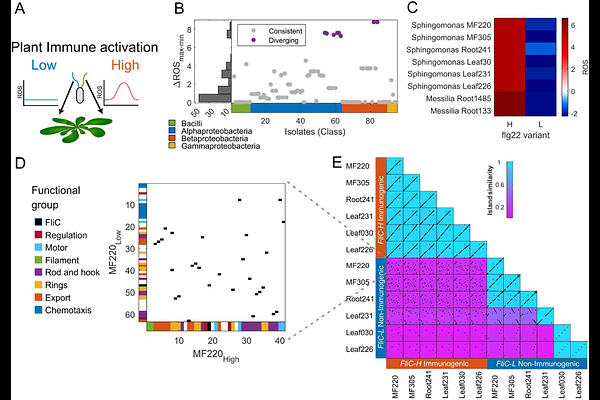
AbstractPlants monitor their environment for microbial invaders using pattern-recognition receptors that detect microbe-associated molecular patterns (MAMPs). Flagellin, the main component of bacterial flagellum, contains the flg22 epitope recognized by the plant immune receptor FLS2. Immune recognition can create an evolutionary conflict, requiring bacteria to balance flagellar function and immune evasion. Here, we show that plant-associated Sphingomonads resolve this constraint by partitioning two flagellar functions, motility and colonization, across two divergent and independently acquired flagellin genes. Comparative genomics revealed widespread coexistence of FliC proteins expressing either an immunogenic variant (FliC-H) or a non-immunogenic variant (FliC-L). The non-immunogenic FliC-L is necessary and sufficient for full directional swimming, whereas FliC-H is dispensable for swimming, but sufficient for full attachment and colonization. Flagellin expression patterns mirror these functions. Thus, FLS2 recognizes the flagellar variant required for colonization rather than motility, potentially restricting colonizing bacteria from entering internal leaf and root tissues.