SCOPE: Localizing fate-decision states and their regulatory drivers in single-cell differentiation
SCOPE: Localizing fate-decision states and their regulatory drivers in single-cell differentiation
Zhao, Y.; Finkbeiner, C.; Setty, M.; Lin, K.
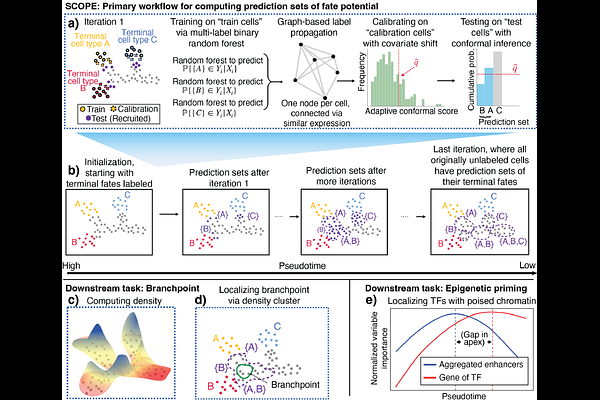
AbstractIdentifying the precise transcriptomic states at which cells commit to a lineage (branchpoints) and the temporal lag in which chromatin accessibility foreshadows gene expression (epigenetic priming) remain fundamental challenges in developmental biology. While current methods for single-cell sequencing data effectively capture developmental flow, they often lack a principled mechanism for delineating the discrete boundaries, a crucial aspect required to map the molecular logic of lineage commitment. We present SCOPE (Semi-supervised Conformal Pre- diction), a framework that transforms high-dimensional single-cell measurements into rigorous, discrete prediction sets of all plausible future fates. By formalizing fate uncertainty via conformal inference, SCOPE localizes the precise biological windows during which multipotent progenitors specify their fate. In multi-omic data, SCOPE uncovers epigenetic priming and identifies its driving transcription factors by detecting regimes where chromatin-derived prediction sets resolve toward terminal fates significantly before their transcriptomic counterparts. We apply SCOPE across simulations, lineage-traced mouse hematopoiesis, multiple human hematopoietic datasets, and human retinogenesis to demonstrate its broad applicability and ability to recapitulate known fate specification drivers. Ultimately, SCOPE provides a statistically grounded foundation for localizing fate decisions across biological replicates and modalities, offering a robust tool for identifying the onset of lineage specification in complex developmental systems.