StrucTTY: An Interactive, Terminal-Native Protein Structure Viewer
StrucTTY: An Interactive, Terminal-Native Protein Structure Viewer
Jang, L. S.-e.; Cha, S.; Steinegger, M.
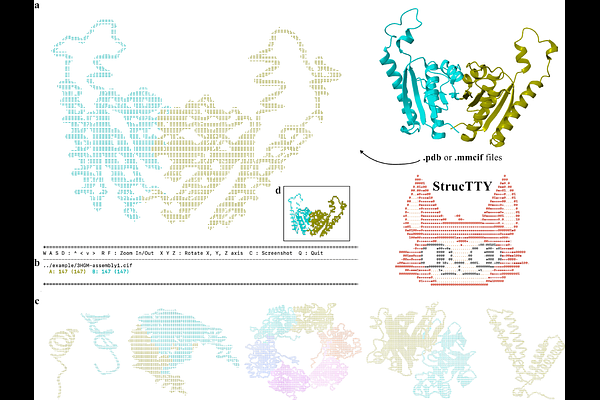
AbstractTerminal-based workflows are central to large-scale structural biology, particularly in high-performance computing (HPC) environments and SSH sessions. Yet no existing tool enables real-time, interactive visualization of protein backbone structures directly within a text-only terminal. To address this gap, we present StrucTTY, a fully interactive, terminal-native protein structure viewer. StrucTTY is a single self-contained executable that loads mulitple PDB and mmCIF files, normalizes three-dimensional coordinates, and renders protein structures as ASCII graphics. Users can rotate, translate, and zoom in on structures, adjust visualization modes, inspect chain-level features and view secondary structure assignments. The tool supports simultaneous visualization of up to nine protein structures and can directly display structural alignments using Foldseek's output, enabling rapid comparative analysis in headless environments. The source code is available at https://github.com/steineggerlab/StrucTTY