Benchmarking three simple DNA staining-based image metrics for live-cell tracking of chromatin organization
Benchmarking three simple DNA staining-based image metrics for live-cell tracking of chromatin organization
Kang, M.; Cabral, A. T.; Sawant, M.; Thiam, H. R.
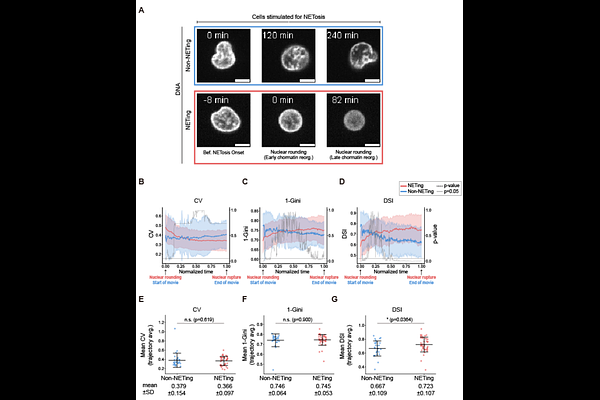
AbstractQuantifying chromatin-state dynamics in living cells remains challenging, in part because most methods require fixation or cell lysis. Here, we benchmark and introduce three simple live-cell image-derived metrics computed from routine DNA staining - the coefficient of variation (CV), 1-Gini, and the Diffuse Signal Index (DSI), introduced here - as fixation-free readouts of chromatin state. Using HL60-derived neutrophils (dHL-60) undergoing NETosis as a model system with a pronounced compact-to-decompact chromatin transition, we show that all three metrics track progressive chromatin reorganization in live-cell trajectories, but differ markedly in sensitivity: DSI provides the strongest trajectory-level discrimination between NETing and non-NETing cells, followed by 1-Gini and CV. Comparison with Tn5-based chromatin accessibility measurements in fixed cells further shows that all three metrics correlate with chromatin accessibility, supporting their biological relevance. Together, our results provide a practical framework for extracting chromatin-state readouts from routine live-cell DNA staining and identify DSI as the most discriminative metric for tracking chromatin reorganization in this benchmark.