Exploring the Energy Landscape of Hairpin Folding using the TIS-DNA model
Exploring the Energy Landscape of Hairpin Folding using the TIS-DNA model
Baratam, K.; Chakraborty, D.
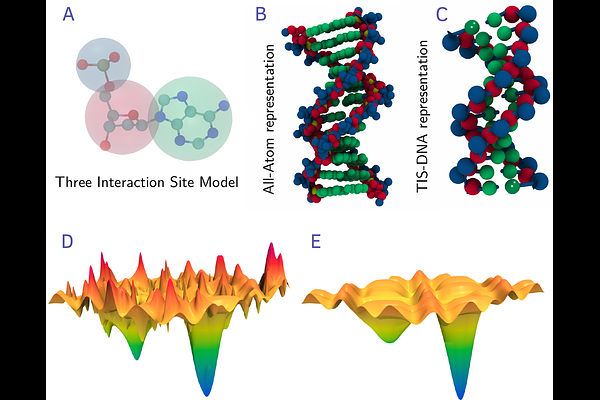
AbstractCoarse-grained (CG) modeling of DNA has been gathering steam in recent years due to the limitations imposed by current computer hardware, and deficiencies of atomistic force-fields. Specifically, CG simulations have emerged as a potent tool for exploring complex energy landscapes underlying biochemical processes, such as DNA transcription, as well as material design based on programmable self-assembly. In this chapter, we illustrate how the Three Interaction Site (TIS) model for DNA, a robust coarse-graining framework, can be used to study the folding landscape of DNA hairpins. We show that despite its simplicity, the TIS-DNA model quantitatively describes the hairpin folding thermodynamics and recapitu- lates many features of the kinetics, including the multiplicity of pathways. The free energy landscape exhibits 'single-funnel' character with a distinct bias towards the folded state. It is likely that folding initiates through non-specific collapse of the DNA chain, involving multiple excursions on the energy landscape, until the opposing strands are approximately aligned. Subsequently, the loop region becomes more ordered, and after the first native- contact nucleates, the rest of the process becomes essentially downhill.