vToxiNet: a biologically constrained deep learning framework for interpretable prediction of drug-induced hepatotoxicity
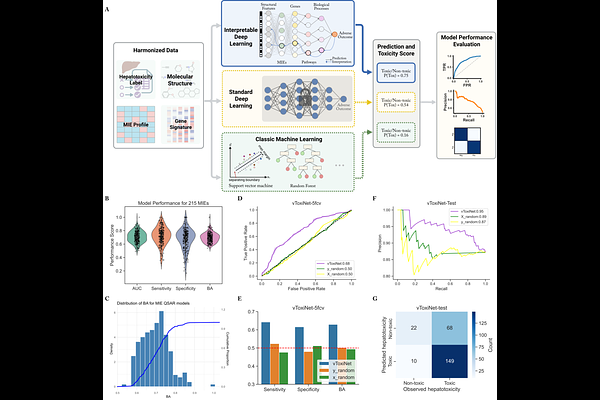
vToxiNet: a biologically constrained deep learning framework for interpretable prediction of drug-induced hepatotoxicity
Jia, X.; Wang, T.; Russo, D. P.; Aleksunes, L. M.; Xiao, S.; Zhu, H.
AbstractHepatotoxicity remains a leading cause of drug attrition and post-marketing withdrawal, resulting from diverse and complex toxicity mechanisms. Traditional in vitro models can only capture a limited subset of toxicity pathways, and animal studies face translational and ethical limitations. Regulatory agencies have therefore promoted new approach methodologies, including human-relevant assays, omics technologies, and computational models to improve predictive toxicology and support evidence-based decision-making. However, most machine learning models for hepatotoxicity either rely solely on chemical structure or operate as black boxes, limiting mechanistic interpretability and broader applicability. Here, we introduce the virtual toxicity network (vToxiNet), a biologically constrained deep learning framework that embeds systems toxicology knowledge directly into neural network architecture for interpretable hepatotoxicity prediction. vToxiNet integrates chemical descriptors, high-throughput assay responses, transcriptomic signatures, and Reactome pathway hierarchy to construct a virtual adverse outcome pathway network. Across cross-validation and multiple external validation datasets, vToxiNet demonstrates robust predictive performance and generalizes to previously unseen chemicals. Importantly, interpretation of vToxiNet enables gene and pathway-level attribution, supporting mechanism-informed hazard characterization and chemical prioritization. These results demonstrate that encoding biological hierarchy as architectural constraints enables both predictive accuracy and mechanistic insight, establishing a generalizable framework for modeling complex biological outcomes.