Automated mini-bioreactors reveal the temporal dynamics and multi-omics responses of CRISPRi knockdowns in Pseudomonas putida
Automated mini-bioreactors reveal the temporal dynamics and multi-omics responses of CRISPRi knockdowns in Pseudomonas putida
Saavedra, M. A.; Grassi, S.; Jespersen, M. G.; Rocha, C.; Kandasamy, V.; Nikel, P. I.; Nielsen, L. K.; Donati, S.
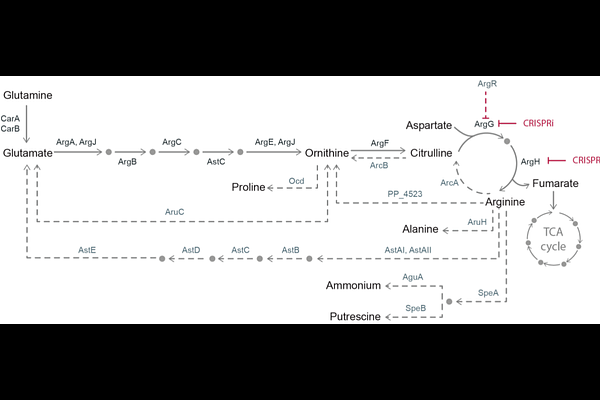
AbstractCharacterizing CRISPR interference (CRISPRi) phenotypes presents a fundamental temporal challenge: pre-existing overabundance of target proteins can mask early silencing, requiring extended growth for dilution, yet prolonged repression rapidly selects for escaper mutants. To resolve this, we integrated a tightly regulated CRISPRi system in Pseudomonas putida with an automated mini-bioreactor platform operating in turbidostat mode. By maintaining continuous exponential growth, we mapped the exact temporal dynamics of essential gene silencing. We identified a critical observation window between 17 and 27 hours (7-9.5 cell doublings) where repression exerts its maximum physiological impact, directly preceding population takeover by target-site mutated escapers. Applying this workflow to the arginine biosynthesis pathway, multi-omics profiling disentangled transient physiological buffering from long-term mutational events, and revealing that argH and argG knockdowns trigger highly diverse metabolomic perturbations. This scalable framework overcomes batch culture limitations, ensuring precise temporal control for accurate phenotypic characterization and reliable functional genomics.