Directional co-transcriptional folding and pausing create kinetic checkpoints for riboswitch-controlled gene expression
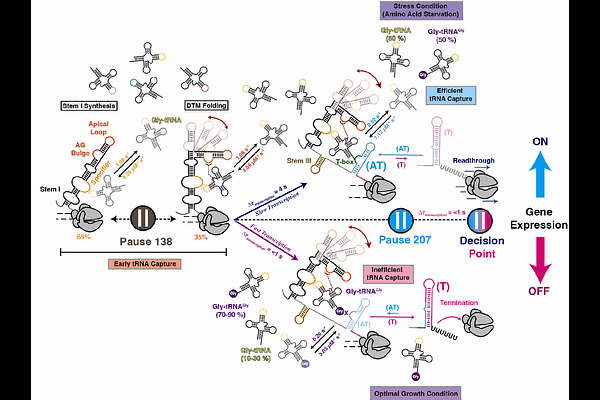
Directional co-transcriptional folding and pausing create kinetic checkpoints for riboswitch-controlled gene expression
Chauvier, A.; Cabello-Villegas, J.; Nikonowicz, E.; Walter, N. G.
AbstractTranscriptional riboswitches typically regulate gene expression by sensing small metabolites or ions that cannot be fluorophore-labeled without perturbing recognition, limiting direct observation of ligand binding during transcription. The GlyQS T-box riboswitch instead senses the uncharged 3' end of a macromolecular ligand, tRNAGly, enabling visualization of co-transcriptional ligand binding. Using single-molecule fluorescence microscopy, we show that tRNAGly dynamically samples discrete structural domains of the nascent riboswitch as transcription proceeds. Productive recognition depends on a hierarchical 5'-to-3' folding pathway shaped by transcription rate and pausing. Pausing enhances tRNA anchoring, stabilizes the riboswitch-tRNA complex, and selectively promotes RNA polymerase readthrough when an uncharged 3' end is presented. These results support a kinetic checkpoint model in which early recruitment increases local ligand concentration, while pause-mediated stabilization converts transient encounters into committed regulatory decisions. More broadly, our findings identify directional co-transcriptional folding and transcriptional pausing as principles synergistically enabling selective and robust riboswitch function.