System-Wide Proteomic Remodeling in Spinal Muscular Atrophy Reveals Tissue-Specific Responses and Partial Rescue by SMN Restoration
System-Wide Proteomic Remodeling in Spinal Muscular Atrophy Reveals Tissue-Specific Responses and Partial Rescue by SMN Restoration
Vrettou, S.; Mueller, S.; Wirth, B.
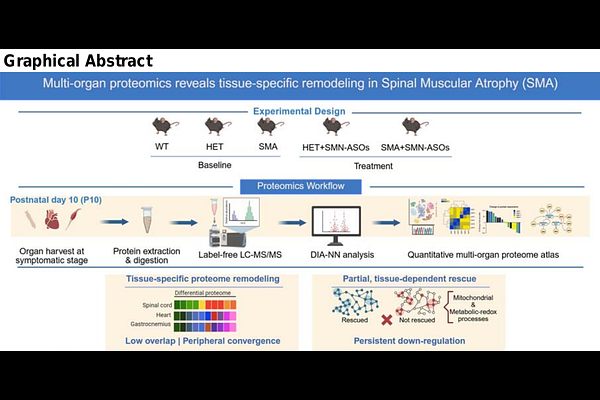
AbstractSpinal muscular atrophy (SMA), traditionally defined as a neuromuscular disorder characterized by degeneration of lower motor neurons, is increasingly recognized as a multi-organ disease. SMA is caused by deficiency of the survival motor neuron (SMN) protein below a critical threshold required for cellular homeostasis. While motor neurons are particularly vulnerable, the ubiquitous expression and fundamental functions of SMN result in widespread perturbations across multiple tissues. Here, we generated a label-free quantitative proteomics atlas of spinal cord, heart, and gastrocnemius muscle from wild-type, heterozygous, and SMA mice at the symptomatic stage, including cohorts treated, at postnatal day 1 (P1), with a systemic suboptimal dose of SMN antisense oligonucleotides (SMN-ASOs), resulting in partial SMN restoration. SMN deficiency induced pronounced, tissue-specific proteome remodeling, with peripheral tissues exhibiting broader molecular alterations than spinal cord. Cross-tissue analyses revealed limited overlap, although heart and muscle showed partial convergence in metabolic and mitochondrial-associated pathways. SMN-ASO treatment partially repositioned these proteomes toward control states; however, restoration was incomplete and strongly tissue-dependent, with persistent dysregulation of mitochondrial and metabolic pathways. These findings demonstrate that SMN deficiency drives systemic yet heterogeneous proteome remodeling and that partial SMN restoration does not fully reverse established molecular alterations.