GeNETop: Context-Specific Genome-Scale Constrained Models Using Network Topology, Flux Variability, and Transcriptomics
GeNETop: Context-Specific Genome-Scale Constrained Models Using Network Topology, Flux Variability, and Transcriptomics
Troitino-Jordedo, D.; Mansouri, A.; Minebois, R.; Querol, A.; Remondini, D.; Balsa-Canto, E.
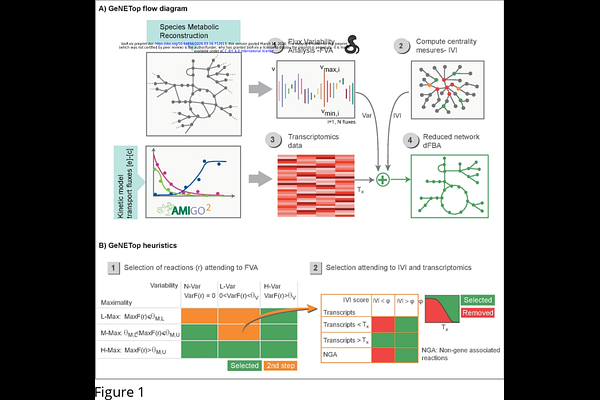
AbstractContext-specific genome-scale metabolic models are critical tools for studying cellular metabolism under dynamic conditions. However, most existing methods for deriving these models are designed for steady-state settings and may fail to preserve reactions required for transient metabolic shifts, thereby limiting their compatibility with dynamic FBA. Here, we present GeNETop, a methodology for deriving context-specific GEMs designed to preserve dynamic compatibility. GeNETop integrates flux variability analysis (FVA), network topology metrics based on the Integrated Value of Influence (IVI), and transcriptomic data to identify reactions that are both flux-flexible and structurally influential. Reactions are prioritized based on variability and maximality indices, while topology and gene expression guide further refinement, reducing dependence on fixed expression thresholds. Using batch fermentation of Saccharomyces cerevisiae as a case study, we evaluate GeNETop against established methods for context-specific metabolic reconstruction. The resulting networks remain dynamically feasible across growth phases, capture key metabolic transitions, reduce non-essential reactions, and maintain computational tractability. Overall, GeNETop enables context-specific metabolic reconstructions that are compatible with dynamic simulations while maintaining computational efficiency. By overcoming key limitations of existing approaches, the method supports a more accurate representation of time-dependent metabolic processes in biotechnology and systems biology.