A run-length-compressed skiplist data structure for dynamic GBWTs supports time and space efficient pangenome operations over syncmers
A run-length-compressed skiplist data structure for dynamic GBWTs supports time and space efficient pangenome operations over syncmers
Durbin, R.
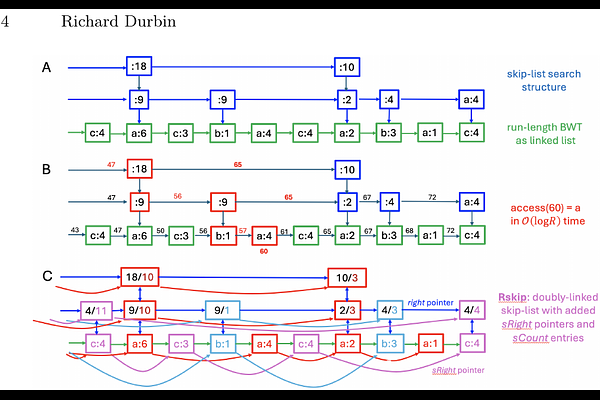
AbstractSkiplists (Pugh, 1990) are probabilistic data structures over ordered lists supporting O(log N) insertion and search, which share many properties with balanced binary trees. Previously we introduced the graph Burrows-Wheeler transform (GBWT) to support efficient search over pangenome path sets, but current implementations are static and cumbersome to build and use. Here we introduce two doubly-linked skiplist variants over run-length-compressed BWTs that support O(log N) rank, access and insert operations. We use these to store and search over paths through a syncmer graph built from Edgar's closed syncmers, equivalent to a sparse de Bruijn graph. Code is available in rskip.[ch] within the syng package at github.com/richarddurbin/syng. This builds a 5.8 GB lossless GBWT representation of 92 full human genomes (280Gbp including all centromeres and other repeats) single-threaded in 52 minutes, on top of a 4GB 63bp syncmer set built in 37 minutes. Arbitrarily long maximal exact matches (MEMs) can then be found as seeds for sequence matches to the graph at a search rate of approximately 1Gbp per 10 seconds per thread.