A genome-wide atlas of meiotic recombination intermediates reveals distinct modes of DNA repair that direct crossovers away from transcriptionally marked genes
A genome-wide atlas of meiotic recombination intermediates reveals distinct modes of DNA repair that direct crossovers away from transcriptionally marked genes
Henfrey, C.; Print, E.; Zhang, G.; Hinch, R.; Maudlin, I.; Moralli, D.; Davies, B.; Donnelly, P.; Hinch, A. G.
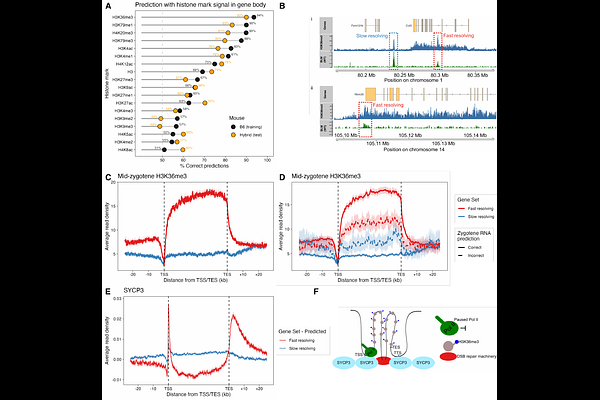
AbstractCrossovers are essential for accurate chromosome segregation in meiosis. Yet the programmed DNA double-strand breaks that initiate them frequently occur in genes and pose a risk to transcription required for gametogenesis. How meiotic cells reconcile these competing demands has remained unclear. Here, we generate a genome-wide in vivo atlas of meiotic recombination intermediates across ~42,000 hotspots by mapping repair proteins BLM, HFM1, and RPA in wild-type and genome-engineered mutant mouse testes. These maps reveal two distinct modes of break repair: a fast-resolving class with short-lived intermediates that are repaired predominantly as non-crossovers, and a slower class with persistent intermediates that give rise to nearly all crossovers. Fast-resolving hotspots occur almost exclusively within a deeply conserved set of ~4,500 genes marked by structural and chromatin features established during an early stage of meiotic transcription. This transcriptional memory predicts repair fate with high accuracy across mouse subspecies and sexes. Across widely diverged mammals, including humans and cattle, orthologous genes show similar crossover suppression. Our findings reveal an early bifurcation between crossover and non-crossover repair that is governed by the transcriptional context of meiotic breaks. Together, they establish an evolutionarily conserved principle in which crossovers are directed away from transcriptionally important genes, thereby safeguarding gene function and shaping their evolution.