Contrasting transcriptional responses and genetic determinants underlie Zymoseptoria tritici adaptation mechanisms to simulated host defense environments
Contrasting transcriptional responses and genetic determinants underlie Zymoseptoria tritici adaptation mechanisms to simulated host defense environments
Minana-Posada, S.; Feurtey, A.; McDonald, B. A.; Lorrain, C.
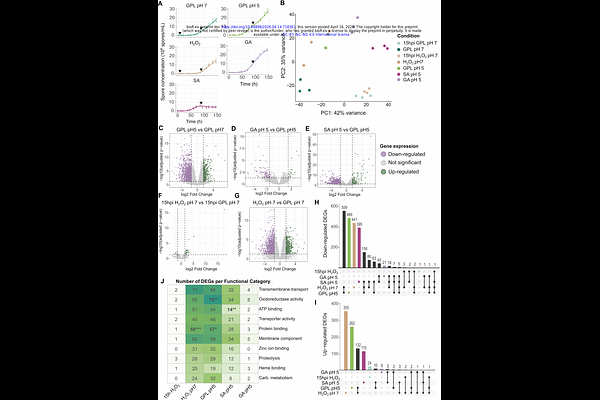
AbstractSuccessful colonization of the wheat apoplast requires that Zymoseptoria tritici tolerate host-derived stresses, but the mechanisms underlying this adaptation remain poorly understood. We combined phenotypic assays, transcriptomics, and genome-wide association analyses to characterize fungal responses to acidic pH, salicylic acid, gibberellic acid, and oxidative stress. Exposure to salicylic acid inhibited in vitro growth across a global collection of 411 Z. tritici strains, whereas acidic pH promoted growth, illustrating contrasting effects on pathogen performance of environments simulating host-defense responses. At the transcriptional level, acidic pH and oxidative stress induced the strongest and most similar responses, while salicylic acid elicited a more distinct transcriptional program and gibberellic acid caused only limited transcriptional changes. Although the sets of differentially expressed genes were largely condition specific, overlapping enrichment of transport- and redox-related functions across conditions indicated shared transcriptional responses. K-mer based genome-wide association mapping identified five candidate loci associated with growth under acidic pH, gibberellic acid and salicylic acid, including four loci specific to a single growth condition. These loci colocalized with genes implicated in cell wall remodeling, nitrogen metabolite regulation, proteostasis, and ubiquitin-related processes. This study highlights the multifaceted strategies employed by Z. tritici to navigate environments simulating host-defense responses, involving shared and environment-specific adaptations. We provide new insights into the genetic and molecular basis of fungal resilience, with implications for understanding pathogen-host interactions.