Explainable AI for end-to-end pathogen target discovery and molecular design
Explainable AI for end-to-end pathogen target discovery and molecular design
Polonio, A.; Perez-Garcia, A.; Fernandez-Ortuno, D.; Jimenez-Castro, L.
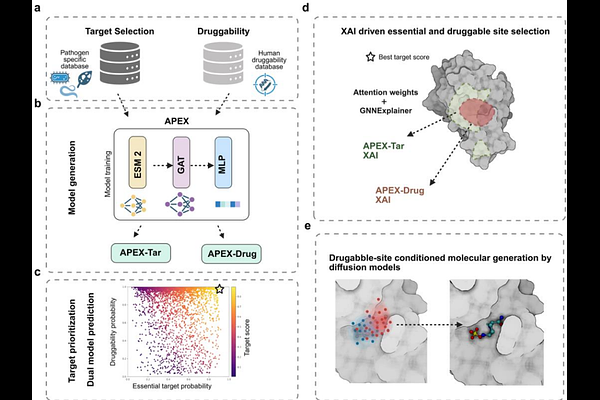
AbstractDrug discovery is often constrained by target identification, a bottleneck especially acute in antimicrobial development and the fight against emerging fungicide resistance. We present APEX (Attention-based Protein EXplainer), an explainable AI framework for cross-species, proteome-scale target discovery and pocket-guided molecular design. APEX combines ESM-2 evolutionary embeddings, graph attention networks, and a multilayer perceptron to train pathogen-specific essentiality and virulence predictors (APEX-Tar) alonsgside a universal druggability model (APEX-Drug). Attention maps and GNNExplainer-derived subgraphs highlight residues and pockets driving predictions, enabling direct conditioning of structure-based diffusion models for inhibitor generation. APEX-Tar recovers known fungal targets (endopolygalacturonase 1, Hog1 MAPK) and proposes new candidates, including fungal GmrSD and bacterial YadV. APEX-Drug recapitulates established fungicide sites ({beta}-tubulin, cytochrome b), guides putative inhibitor design for GmrSD, and identifies in YadV a previously undescribed pocket distinct from known pilicide sites. Together, APEX offers a kingdom-agnostic pipeline for explainable target prioritization and guided molecular design.