Mutation-induced reshaping of protein conformational dynamics revealed by a coarse-grained modeling framework
Mutation-induced reshaping of protein conformational dynamics revealed by a coarse-grained modeling framework
Lee, B. H.; Scaramozzino, D.; Piticchio, S.; Orellana, L.
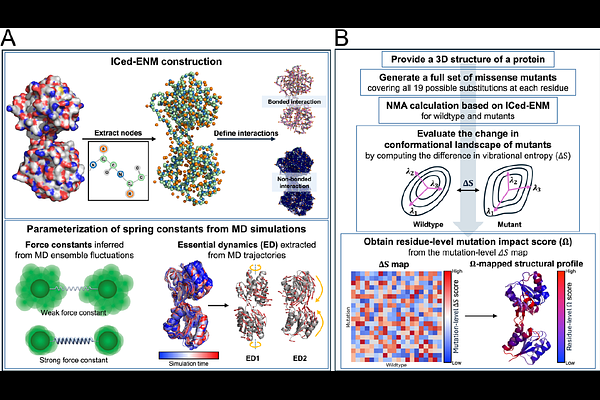
AbstractDisease-related missense mutations reshape protein conformational energy landscapes, thereby altering biological function. However, mechanistically linking sequence variation to changes in conformational dynamics remains challenging for both experimental and computational approaches. Here, we introduce an internal-coordinate-based, essential-dynamics-refined elastic network model (ICed-ENM) that improves the physical fidelity of normal modes while capturing subtle mutation-induced side-chain effects and preserving computational efficiency. By constraining bond-length and bond-angle fluctuations and refining mode subspaces against experimentally observed collective motions, ICed-ENM provides a stable, structure-encoded description of intrinsic protein dynamics. Building on this framework, we developed a systematic mutation-scanning analysis that quantifies mutation impact as changes in vibrational entropy, providing a dynamic measure of mutation-induced redistribution within conformational energy landscapes. Validation against all-atom molecular dynamics simulations demonstrates that residues predicted as mutation hot spots induce substantial reshaping of free-energy landscapes, consistent with altered intrinsic conformational dynamics. Extending this analysis across a curated protein structure dataset reveals global patterns of mutation sensitivity across diverse structural and physicochemical contexts. Notably, these trends align with large-scale public mutation datasets, suggesting that our framework captures features relevant to pathogenic variation. Together, ICed-ENM and the associated mutation-scanning pipeline provide a scalable and mechanistically interpretable strategy to identify mutation-sensitive regions and substitutions, offering deeper insight into how sequence variation reshapes functional conformational landscapes.