PHENOCAUZ: Linking Human Symptoms, Drug Side Effects and Efficacy to Their Molecular Causes Using Mendelian Disease Biology
PHENOCAUZ: Linking Human Symptoms, Drug Side Effects and Efficacy to Their Molecular Causes Using Mendelian Disease Biology
Zhou, H.; Skolnick, J.
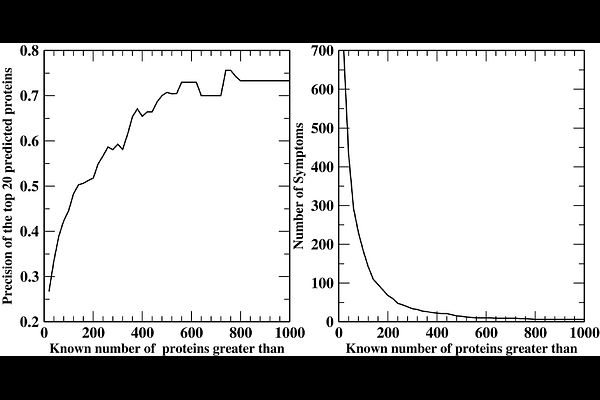
AbstractHuman diseases and adverse drug reactions are ultimately recognized through clinical symptoms, yet the molecular determinants of most symptoms remain unknown. To address this key issue, we present PHENOCAUZ, a computational framework that links symptoms to their causative proteins by integrating Mendelian phenotype - gene relationships with molecular features of proteins. Starting from symptom annotations derived from Mendelian phenotypes and their causal genes, PHENOCAUZ identifies biological pathways and processes associated with individual symptoms and trains a machine learning model to predict symptom-causing proteins beyond those currently implicated in Mendelian diseases. The approach is motivated by the hypothesis that if dysfunction of a protein produces a symptom in a Mendelian disorder, the same protein may contribute to the same symptom in a complex disease or cause drug toxicity. Benchmarking across 2,344 symptoms and 4,828 Mendelian proteins using leave-one-out cross-validation yielded an estimated precision of approximately 0.70 among the top predictions. Predicted symptom-protein relationships show strong pathway-level agreement with literature-curated symptom-protein associations, efficacious drug targets and disease mode-of-action proteins. PHENOCAUZ also enables practical applications including prediction of severe drug side effects and identification of candidate therapeutics for ovarian, prostate, and breast cancers as well as noncancer diseases such as dementia and Crohn's disease. These results demonstrate that Mendelian disease biology provides a powerful route to connect clinical symptoms with their molecular determinants and translate those insights into drug discovery and safety prediction.