The genome of the Delisea pulchra: a resource for the study of chemical host-microbe interactions in red algae
The genome of the Delisea pulchra: a resource for the study of chemical host-microbe interactions in red algae
Dittami, S. M.; Hudson, J.; Brillet-Gueguen, L.; Ficko-Blean, E.; Tanguy, G.; Rousvoal, S.; Legeay, E.; Markov, G. V.; Delage, L.; Godfroy, O.; Corre, E.; Collen, J.; Leblanc, C.; Egan, S.
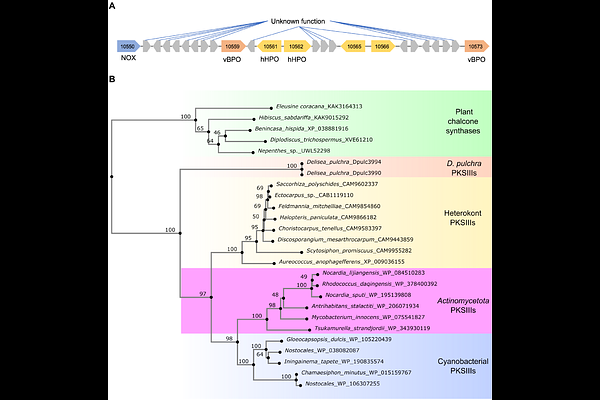
AbstractBackground: Red macroalgae (Rhodophyta) are ecologically and economically important marine primary producers, yet genomic resources for most species remain scarce. Delisea pulchra, a temperate red alga known for its halogenated furanone-based chemical defenses, serves as a model for studying algal-microbe interactions, antifouling mechanisms, and disease dynamics. Results: Here we present a high-quality genome assembly of this species. The nuclear genome comprises 134 Mbp across 271 contigs with an N50 of 1.47 Mbp and encodes 13,387 predicted protein-coding genes. Comparative genomics with other red algae revealed expansions in gene families involved in DNA methylation, and oxidative stress responses, including glutathione S-transferases and superoxide dismutases. Analysis of glycosyltransferases, sulfatases, and sulfurylases implicated in galactan biosynthesis suggests D. pulchra possesses a complex and potentially novel extracellular matrix. We also identified several vanadium haloperoxidases (vHPOs), heme-dependent haloperoxidases (hHPOs), and two type III polyketide synthase (PKS) genes unique to the D. pulchra, which together represent promising candidate genes for bromofuranone production. Conclusion: The D. pulchra genome provides a foundation for molecular investigations into defense, signaling, and host-microbe interactions. It has been deposited at the European Nucleotide Archive under accession number PRJEB101077. All datasets, annotations, and interactive tools for exploring the genome are also available through the Rhodoexplorer portal at https://rhodoexplorer.sb-roscoff.fr.