An epigenetic bifunctional that toggles between transactivation and repression
An epigenetic bifunctional that toggles between transactivation and repression
Sadagopan, A.; Carson, M.; Zamurs, E. J.; Srikonda, S.; Weiss, C. N.; Bond, M. J.; Sodhi, A.; Donovan, K. A.; Ryan, J. K.; Fischer, E. S.; Stegmaier, K. S.; Viswanathan, S. R.; Ebert, B. R.; Gibson, W. J.
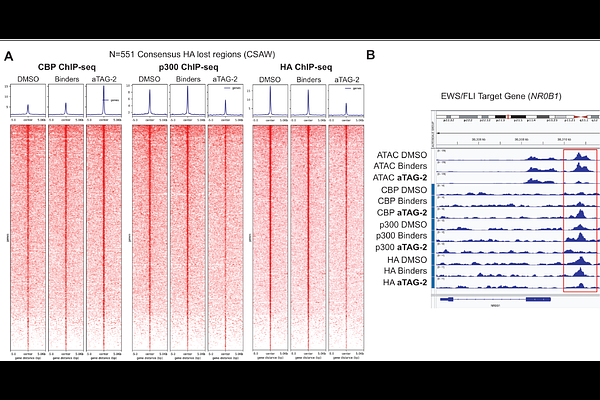
AbstractThe targeted modulation of gene expression with bifunctional small molecules enables the precise control of cellular and biological processes. To screen for ligands that could be used to induce gene expression, we conjugated the high affinity FKBP(F36V) binder, AP1867, to known high-affinity binders of activating epigenetic machinery. We tested these bifunctionals in a FKBP(F36V)-tagged transcription factor reporter system and found bifunctional induced transactivation is relatively common, being observed for bifunctionals with BET ligand JQ1, p300/CBP ligand GNE-781, CDK9 ligand SNS-032, and BRD9 ligand iBRD9. aTAG-2 (mAP1867-C8-GNE781) was identified as the strongest and most potent transactivator, possessing single-digit nanomolar activity. When tested in models where oncogenic RNA binding protein-transcription factor fusion proteins have been FKBP(F36V)-tagged, we unexpectedly observed rapid collapse of the fusion transcriptional program. In a tagged Ewing sarcoma model, aTAG-2 exhibits at least three distinct mechanisms of action: i) RIPTAC mediated p300/CBP inhibition, ii) ubiquitination- and ternary complex-dependent EWS/FLI degradation, and iii) replacement of p300 with CBP at EWS/FLI bound chromatin loci. Together, these data establish bifunctionals targeting p300/CBP that toggle between a program of ultra-potent transactivation and repression depending on cellular context. Overall demonstrating that induced proximity with a given ligand does not encode a fixed functional outcome.