Scalable genotyping in fixed transcriptomes resolves clonal heterogeneity via single-cell sequencing
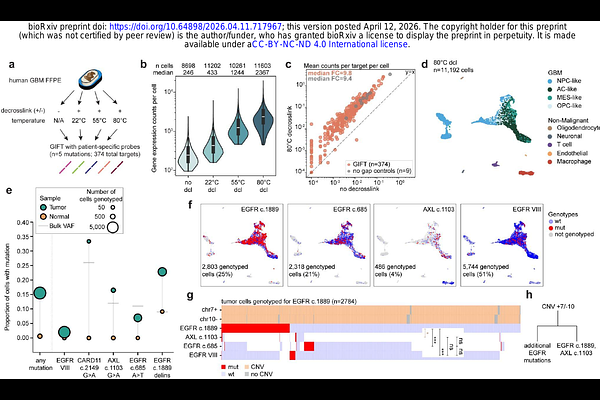
Scalable genotyping in fixed transcriptomes resolves clonal heterogeneity via single-cell sequencing
Blattman, S. B.; Maslah, N.; Varela, A. A.; Kumpaitis, K.; Nalbant, B.; Snopkowski, C.; Mariani, M.; Kida, L. C.; Takizawa, M.; Ratnayeke, N.; Yu, K. K. H.; Fernandes, S.; Mousavi, N.; Borgstrom, E.; Vallejo, D.; Boghospor, L.; Xin, R.; Mignardi, M.; Wu, S.; Scarlott, N.; Delgado-Rivera, L.; Kumar, P.; Krishnan, S.; Giraudier, S.; Kiladjian, J.-J.; Howitt, B. E.; Kohlway, A.; Lund, P.; Pe'er, D.; Chaligne, R.; Lareau, C. A.
AbstractSingle-cell transcriptomics has revolutionized our understanding of heterogeneous cell populations. However, technical limitations of widely-used platforms have limited our ability to link transcriptional states to somatic mutations within the same cells at scale. Here, we introduce Genotyping in Fixed Transcriptomes (GIFT), a novel assay for simultaneous detection of hundreds of targeted genetic variants and whole transcriptome profiles in single cells. The core innovation of GIFT is a rationally designed gapfilling reaction between adjacent single-stranded DNA (ssDNA) probes that barcodes native transcript sequence to enable highly-specific targeted mutation detection. GIFT achieves >99% genotyping accuracy and flexible capture of hundreds of mutations per cell, including in FFPE (Formalin-Fixed Paraffin-Embedded) tissue, enabling clonal lineage tracing in heterogeneous settings. We demonstrate the unique scalability of GIFT by profiling >700,000 cells from 35 donors with myeloproliferative neoplasms (MPN), revealing mutation-dependent hematopoietic responses to systemic inflammation associated with the characteristic JAK2V617 mutation, including an allelic dose gradient of interferon-associated transcriptional programs and transcriptional priming of hematopoietic stem cells that develop into divergent disease states. Together, the unique technical advantages of GIFT enable direct resolution of genotype-to-phenotype relationships via clonal lineage tracing with comprehensive cell state measurements at single-cell resolution.