mRNA stability in response to m6A placement is linked to cell identity in planarians
mRNA stability in response to m6A placement is linked to cell identity in planarians
Hoehn, C.; Pittroff, A.; Gribling-Burrer, A.-S.; Huelsmann, L.; Smyth, R. P.; Kuhn, C. D.
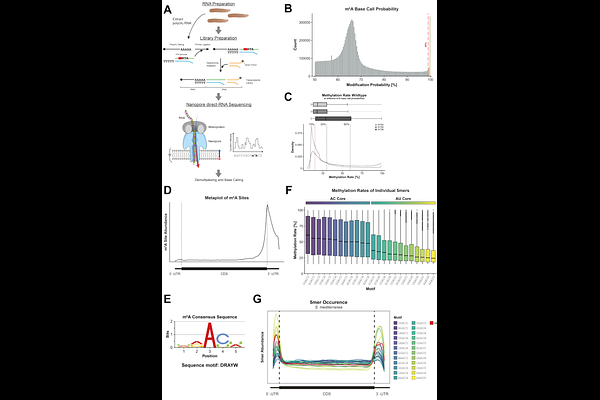
AbstractN6-methyladenosine (m6A) is a widespread internal modification of eukaryotic mRNA that influences transcript fate, including mRNA stability and cell-type-specific gene expression. However, the mechanisms underlying m6A-mediated regulation remain poorly understood in many systems, including the highly regenerative planarian Schmidtea mediterranea. To address this, we generated a high-confidence atlas of ~72,200 m6A sites across the planarian transcriptome using multiplexed direct RNA sequencing. m6A sites follow a DRAYW consensus motif and are highly enriched near stop codons while being largely excluded from coding sequences. This distribution is consistent with an exon length-dependent variant of the exclusion model mediated by the exon junction complex. Knockdown of the m6A writer complex resulted in pronounced, cell-type-specific changes in transcript stability. Destabilized transcripts are enriched for intestinal markers, whereas stabilized transcripts are, amongst others, associated with neoblasts, the adult stem cells of planarians. Transcriptional shut-off experiments further confirmed that m6A has opposing effects on mRNA decay depending on cellular context: it stabilizes transcripts in differentiated cells, while it promotes the degradation of mRNAs associated with neoblasts. Taken together, these results support a model in which cell-type-specific regulation of mRNA stability by m6A plays a crucial role in shaping cell identity in planarians.