Optimizing resource allocation in Miscanthus breeding with sparse testing designs for genomic prediction
Optimizing resource allocation in Miscanthus breeding with sparse testing designs for genomic prediction
Proma, S.; Lubanga, N.; Sacks, E.; Leakey, A. D. B.; Zhao, H.; Ghimire, B. K.; Lipka, A. E.; Njuguna, J. N.; Yu, C. Y.; Seong, E. S.; Yoo, J. H.; Nagano, H.; Anzoua, K. G.; Yamada, T.; Chebukin, P.; Jin, X.; Clark, L. V.; Petersen, K. K.; Peng, J.; Sabitov, A.; Dzyubenko, E.; Dzyubenko, N.; Glowacka, K.; Nascimento, M.; Campana Nascimento, A. C.; Dwiyanti, M. S.; Bagment, L.; Shaik, A.; Garcia-Abadillo, J.; Jarquin, D.
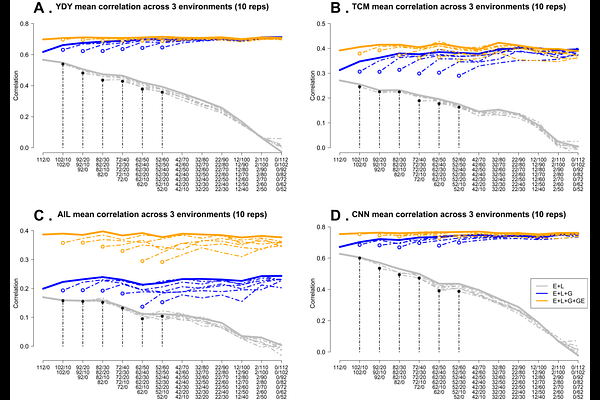
AbstractPhenotyping high-biomass perennial crops is laborious and the rate of genetic gain in perennial crop breeding programs is typically low. So, it is especially important to identify methods that produce efficiency gains in the breeding process. Miscanthus is a C4 perennial grass with favorable characteristics for producing biomass as a feedstock for biofuels and diverse biobased products. Increasing biomass yield will increase profitability and environmental benefits, so is a key target for Miscanthus breeding. In addition, the identification of well-adapted genotypes across a wide range of environmental conditions requires the establishment of multi-environment trials (METs). Sparse testing is a genomic prediction-based strategy that reduces the phenotyping costs in METs by selecting a subset of genotypes to evaluate in a subset of environments and then predicts the performance of the unobserved genotype-environment combinations. A Miscanthus sacchariflorus (MSA) population comprising 336 genotypes observed across three environments was analyzed. Three prediction models considering main effects (environments, genotypes, genomic) and interaction effects (genotype-by-environment; GxE interaction) were implemented for forecasting dry biomass yield (YDY), total culm (TCM), average internode length (AIL), and culm node number (CNN). Multiple calibration sets based on different compositions and sizes were considered to evaluate performance in terms of the predictive ability (PA) and the mean square error (MSE) for a fixed testing set size. The training set size ranged from 52 to 112 to predict a fixed set of 224 unobserved genotypes across all three environments. The results showed that the model accounting for GxE interaction presented the highest PA and the lowest MSE for CNN (PA: ~0.77, MSE: ~0.5) and YDY (PA: ~0.70, MSE: ~1.3) while for TCM and AIL these ranged from ~0.28 to 0.41 and ~1.3 to 4.3, respectively. Overall, varying training sets and allocation strategies did not affect PA and MSE, with 52 non-overlapping and 0 overlapping genotypes per environment as the optimal cost-effective allocation framework. This suggests that implementing sparse testing designs could significantly reduce phenotyping costs by fivefold, without compromising PA in breeding programs for perennial crops such as Miscanthus.