Chiron3D: an interpretable deep learning framework for understanding the DNA code of chromatin looping
Chiron3D: an interpretable deep learning framework for understanding the DNA code of chromatin looping
Hoenig, S.; Grover, A.; Neri, P.; Surdez, D.; Boeva, V.
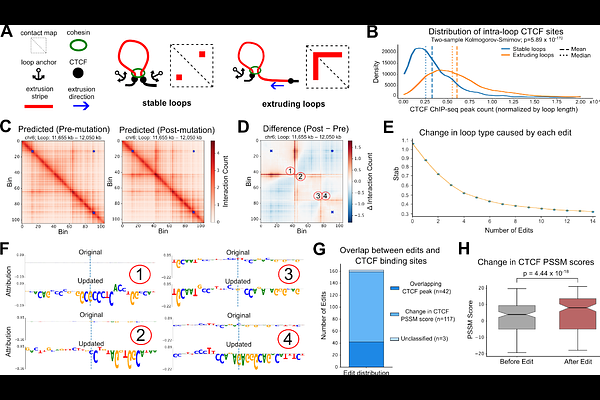
AbstractThree-dimensional folding of the genome into structures such as chromatin loops is essential for gene regulation. Current experimental methods for mapping these structures, like Hi-C and HiChIP, are labor-intensive and require repeated assays to test hypothesized mutation effects. This motivates the need for predictive approaches that reveal the sequence determinants of chromatin loops. In this work, we present a novel and interpretable computational pipeline for predicting CTCF-mediated chromatin loops. We propose Chiron3D, a DNA-only model trained in a cell-type-specific manner to predict CTCF HiChIP contact maps. By leveraging pre-trained embeddings from a foundation model, our approach is competitive with baselines that take CTCF ChIP-seq as additional input, while enabling nucleotide-level attribution to the input DNA sequence. Using our framework, we provide likely mechanistic insights into the physical control of loop dynamics. Specifically, we find that the strength of the loop extrusion anchorage site is largely governed by the amount and binding affinity of CTCF sites at the boundaries. Furthermore, we reveal that loop stability is regulated by the amount of intra-loop CTCF binding sites, where fewer sites within the loop lead to a more stable domain. Using targeted, single-nucleotide edit simulations with Chiron3D, we show that both loop strength and stability can be precisely controlled. Together, these results provide novel mechanistic insights into the physical control of genome organization and highlight the potential of decoding the DNA sequence logic in silico.