Disagreement among variant effect predictors guides experimental prioritization of target proteins
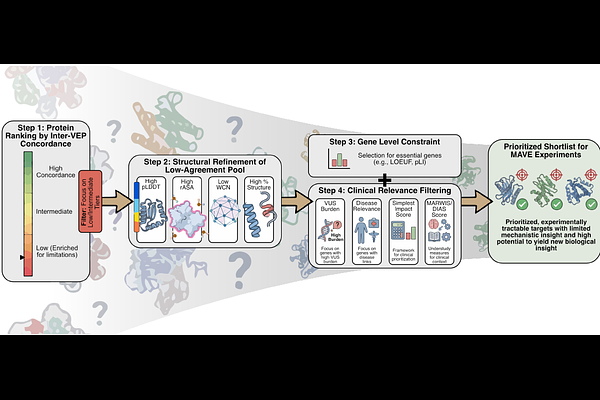
Disagreement among variant effect predictors guides experimental prioritization of target proteins
Jonsson, N. F.; Marsh, J. A.; Lindorff-Larsen, K.
AbstractInterpreting the functional consequences of genetic variation, especially rare missense variants, remains a significant challenge in human genetics. Computational variant effect predictors (VEPs) and multiplexed assays of variant effects (MAVEs) provide complementary approaches, with VEPs offering scalable predictions and MAVEs delivering detailed empirical measurements. However, MAVEs are resource intensive and cannot yet be applied broadly across the proteome, making it important to identify proteins where experimental mapping will be most informative. We hypothesised that MAVEs should be particularly valuable for proteins where computational predictors disagree, as such disagreement may highlight mechanistic blind spots. To test this, we analysed predictions from ten distinct VEPs across more than 13,000 human proteins and quantified inter-predictor concordance. We observed substantial variability across proteins in the degree of agreement across predictors and investigated structural, functional and gene-level features associated with this variation. We find that inter-VEP concordance showed no relationship with agreement to experimental MAVE data. If predictor agreement reflected how intrinsically predictable a protein is, these quantities would be expected to correlate. Their decoupling instead suggests that MAVEs may provide orthogonal information to VEPs, supporting the use of inter-VEP disagreement to prioritise proteins where experimental data will be most informative. We therefore propose using inter-VEP disagreement as a practical strategy to prioritise proteins for experimental characterization. Focusing on proteins with low predictor concordance should maximise the informational value of new MAVEs, and improve variant interpretation in both research and clinical contexts.