Exploration of the structural and functional diversity in the metamorphic RfaH subfamily
Exploration of the structural and functional diversity in the metamorphic RfaH subfamily
Tabilo-Agurto, C.; Gonzalez-Bustos, B.; Reyes, J.; Wang, B.; Palomera, D.; Del Rio-Pinilla, V.; Neira-Mahuzier, C.; Vera-Sandoval, V.; Artsimovitch, I.; Galaz-Davison, P.; Ramirez-Sarmiento, C. A.
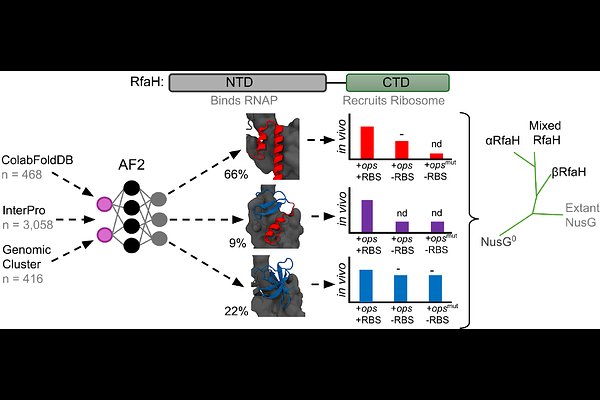
AbstractRfaH, a member of the universally conserved NusG/Spt5 family, activates transcription and translation of virulence genes by bridging the transcribing RNA polymerase to the translating ribosome. Unlike its monomorphic, constitutively active paralog NusG, RfaH fold-switches from an autoinhibited state, wherein its C-terminal domain forms an -helical hairpin bound to the N-terminal domain, into an active NusG-like state through C-terminal domain dissociation and refolding into a {beta}-barrel. RfaH has been proposed to evolve from NusG via gene duplication by first activating genes adjacent to its production site, before developing autoinhibition to selectively control distant genes. Using AlphaFold2, we predicted the structures of thousands of RfaH homologs and identified a subset (~14%) that appear predominantly folded in the active state. In vivo assays confirmed that these putative monomorphic homologs exhibit constitutive activity comparable to known RfaH mutants. Phylogenetic and genomic analyses revealed that these monomorphic proteins form a distinct clade and are preferentially located within or next to long virulence-associated operons. Together, these results further support a stepwise evolutionary model of RfaH specialization through structural transformation.