HERCULES: an integrative deep-learning framework for predicting RNA-binding propensity and mutation effects at single-residue resolution
HERCULES: an integrative deep-learning framework for predicting RNA-binding propensity and mutation effects at single-residue resolution
Fiorentino, J.; Monti, M.; Armaos, A.; Vrachnos, D. M.; Di Rienzo, L.; Tartaglia, G. G.
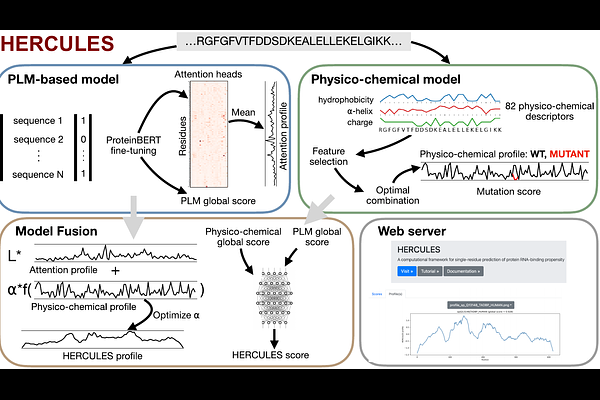
AbstractRNA-binding proteins (RBPs) regulate essential aspects of RNA metabolism, yet accurately identifying RNA-binding domains (RBDs) and quantifying the impact of sequence variation on RNA-binding ability remain challenging. Here, we present HERCULES (Hybrid framEwoRk for RNA-binding domain loCalization and mUtation anaLysis using physicochemical and languagE modelS), a unified sequence-based framework for simultaneous RBD localization, global RNA-binding propensity prediction and mutation effect assessment. HERCULES integrates a fine-tuned protein language model with an explicit residue-level physicochemical module, combining global contextual representations with local mutation-sensitive descriptors. On an independent test set, the HERCULES global score discriminates RBPs from non-RBPs with an AUROC of 0.86. At residue resolution, HERCULES outperforms state-of-the-art sequence-based predictors in identifying canonical, non-canonical and putative RBDs across Pfam-annotated proteins. Using a curated dataset of experimentally validated RNA-binding-disrupting mutations, HERCULES correctly classifies 87% of deleterious variants, including single-amino acid substitutions. Evaluation on experimentally resolved protein-RNA complexes further demonstrates robust residue-level performance and improved generalization when contact annotations are augmented with AlphaFold3-predicted complexes. By unifying domain localization and mutation sensitivity within a single sequence-only framework, HERCULES provides a mechanistically interpretable approach for studying RNA-protein interactions. HERCULES is freely available at https://tools.tartaglialab.com/hercules and as an open-source Python package at https://github.com/tartaglialabIIT/hercules.git.