Conserved protein folds underpin the diversification of secreted proteins in a fungal pathogen
Conserved protein folds underpin the diversification of secreted proteins in a fungal pathogen
Dal'Sasso, T. C. S.; Stukenbrock, E. H.
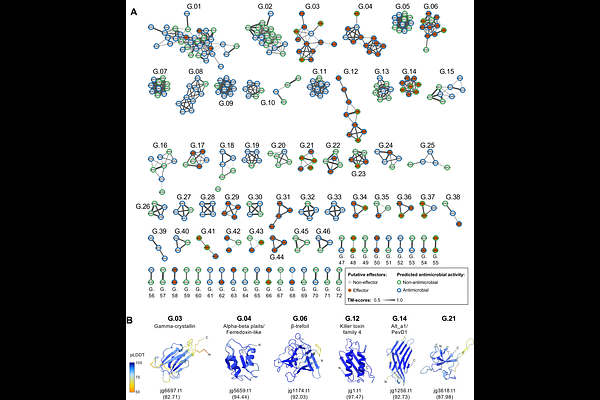
AbstractBACKGROUND During host colonization, fungal plant pathogens secrete effector-like proteins that alter host cell physiology and target plant-associated microbes. However, rapid evolution and low sequence conservation hinder the study and characterization of these proteins. The fungus Zymoseptoria passerinii infects Hordeum spp. and includes lineages adapted to wild and domesticated barley. To date, the evolution of effector-like proteins in this species has not been addressed. RESULTS We combined a set of protein structure-based and network analyses to unravel the secretome of Z. passerinii. We first compared AlphaFold2 and ESMFold predictions to establish the baseline for structural analyses. We identified 72 structural clusters in the secretome, revealing fold-level relationships across divergent sequences. We showed that effector-like proteins with host putative immune-interfering functions evolved from a limited group of protein folds, whereas proteins with predicted antimicrobial properties were distributed across fold groups. Physicochemical comparisons indicate that antimicrobial effectors predominantly emerged through amino acid replacements on common effector-enriched scaffolds in Z. passerinii, reconfiguring surface charge and electrostatics. We analyzed intra- and interspecific structural variation in selected effector-enriched families by comparing Z. passerinii proteins and homologs in the sister species Z. tritici. We describe high structural similarity in core folds, with local variation in loop and surface-exposed regions, consistent with fold stability still enabling functional diversification. CONCLUSIONS The secretome of Z. passerinii is organized around common structural folds that support diverse biological roles, including host manipulation and host-associated microbial interactions. Conserved scaffolds combined with local physicochemical variation likely contribute to rapid adaptive evolution in Z. passerinii.