ECHO: a nanopore sequencing-based workflow for (epi)genetic profiling of the human repeatome
ECHO: a nanopore sequencing-based workflow for (epi)genetic profiling of the human repeatome
Poggiali, B.; Putzeys, L.; Andersen, J. D.; Vidaki, A.
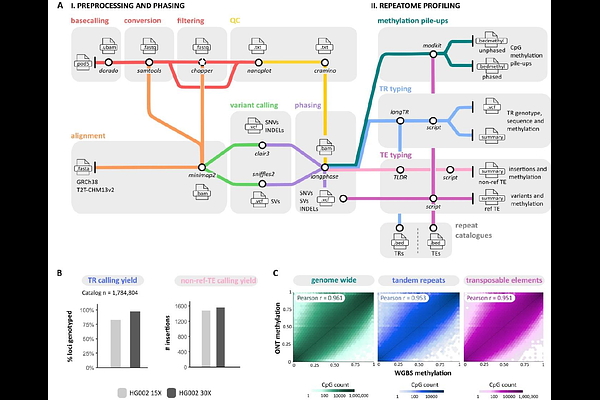
AbstractThe human genome is dominated by repetitive DNA, whose genetic and epigenetic variation plays a key role in gene regulation, genome stability, and disease. Recent advances in long-read sequencing now enable large-scale, haplotype-resolved, and DNA methylation-informative analysis of the human genome, including on previously inaccessible complex and repetitive regions. However, the comprehensive, simultaneous characterisation of the "human repeatome" remains challenging, largely due to the lack of comprehensive tools integrated in a single pipeline that can capture the full spectrum of variation across diverse types of DNA repeats. Here, we present ECHO, a user-friendly, Snakemake-based pipeline for the "(Epi)genomic Characterisation of Human Repetitive Elements using Oxford Nanopore Sequencing". ECHO provides a reproducible and scalable framework for end-to-end analysis of whole-genome nanopore sequencing data, enabling integrative but also tailored (epi)genetic analyses of the human repeatome