Multi-omics definition of the sex-specific glycoproteome of murine tissues
Multi-omics definition of the sex-specific glycoproteome of murine tissues
Kawahara, R.; Hane, M.; Wu, D.; Zhang, B.; Sakamoto, F.; Nakagawa, T.; Omoto, T.; Bienes, K. M.; Bansal, N.; Sumer-Bayraktar, Z.; Chatterjee, S.; Himori, K.; Nagai-Okatani, C.; Kuno, A.; Kashima, M.; Kolarich, D.; Matsui, Y.; Kitajima, K.; Kadomatsu, K.; Sato, C.; Thaysen-Andersen, M.
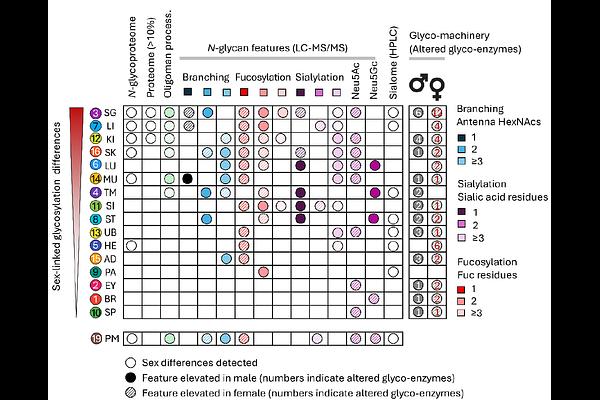
AbstractSex-specific differences in the glycoproteome remain poorly defined despite growing evidence that protein glycosylation is a key determinant of sex biology. Here we present a tissue-resolved glycoproteome atlas of adult male and female C57BL/6J mice, integrating transcriptomics, proteomics and glycoproteomics with sialic acid speciation and lectin microarray profiling across 19 tissues. Quantitative analysis of >26,800 protein- and site-specific N-glycoforms from 1,512 glycoproteins revealed highly distinct tissue glycoproteomes shaped by coordinated regulation of protein abundance and glyco-enzyme expression. Multi-omics integration identified strong glycophenotype-enzyme relationships, including control of tissue sialylation by Cmas and Cmah, suggesting rate-limiting roles in glycosylation. Pronounced sex-linked glycophenotypes were observed in salivary gland, liver and kidney, driven by differences in fucosylation, sialylation and protein abundance, whereas the brain glycome was largely conserved between sexes. An interactive online database (https://igcore.cloud/mta/atlas-viewer/) provides a resource for exploring sex-biased glycosylation across mouse tissues.