Translating Histopathology Foundation Model Embeddings into Cellular and Molecular Features for Clinical Studies
Translating Histopathology Foundation Model Embeddings into Cellular and Molecular Features for Clinical Studies
Cui, S.; Sui, Z.; Li, Z.; Matkowskyj, K. A.; Yu, M.; Grady, W. M.; Sun, W.
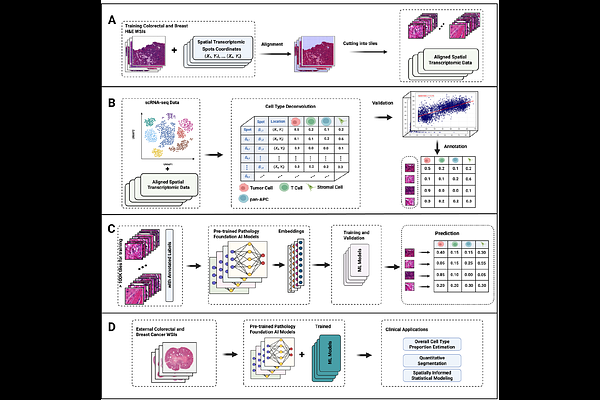
AbstractAI-powered pathology foundation models provide general-purpose representations of histopathological images by encoding image tiles into numerical embeddings. However, these embeddings are not directly interpretable in biological or clinical terms and must be translated into biologically meaningful features, such as cell-type composition or gene expression, to enable downstream clinical applications. To bridge this gap, we developed STpath, a framework that integrates histopathology image embeddings derived from existing pathology foundation models with matched, spatially resolved transcriptomics data. STpath consists of cancer-specific XGBoost models trained to infer cell-type compositions and gene expression from histopathology image tiles. We evaluated STpath in colorectal and breast cancer datasets and showed that it provides accurate estimates of the composition of major cell types and the expression of a subset of genes, with further performance gains achieved by combining embeddings from multiple foundation models. Finally, we demonstrated that STpath inferred features that can be used in downstream studies to evaluate their associations with clinical outcomes.