Uncovering zebrafish embryonic proteome dynamics across 16 time points during the first 24 hours of development
Uncovering zebrafish embryonic proteome dynamics across 16 time points during the first 24 hours of development
Fang, F.; Poulos, W.; Yue, y.; Li, K.; Cibelli, J.; Liu, X.; Sun, L.
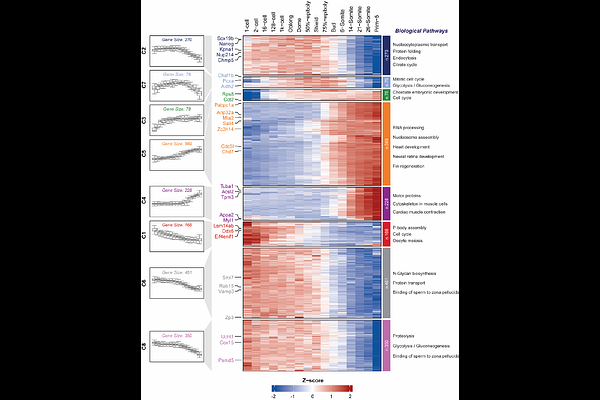
AbstractDefining how proteins change over developmental time is amenable to studies deciphering regulatory genetic networks in vertebrate development, biology, and pharmacology. In an approach toward such quantitative studies of dynamic network behavior, we produced an atlas using the mass spectrometry-based method to investigate protein expression changes across 16 time points from the zygote to the early pharyngula stage zebrafish embryos. We systematically summarize 8 clusters for interrogating changes in protein expression associated with the development of zebrafish embryos. Specifically, we identified a class of zinc finger-related transcription factors primarily located on the long arm of chromosome 4, which are highly expressed during zygotic genome activation. Furthermore, we highlight the power of this analysis to assign developmental stage-specific expression information to chromosomes and tissues. Time-resolved analyses reveal significant discordance between differential transcript and protein expression, whereas no time lag is observed for proteins involved in stable and fundamental biological processes, such as metabolism (e.g., Ppt2a and Gatm), cytoskeletal organization (e.g., Col18a1), and the translation machinery (e.g., Eif4enif1). This atlas offers high-resolution and in-depth molecular insights into zebrafish development, providing a resource for developmental biologists to generate hypotheses for functional analysis of proteins during early vertebrate embryogenesis.