Integrating GlycoSHIELD Modeling and DNA-PAINT SMLM to Map the Glycosylation-Dependent Distri-bution of the Na,K-ATPase
Integrating GlycoSHIELD Modeling and DNA-PAINT SMLM to Map the Glycosylation-Dependent Distri-bution of the Na,K-ATPase
Stojcic, B.; Draczkowski, P.; Patrick, J.; Saeed, M.; Brismar, H.
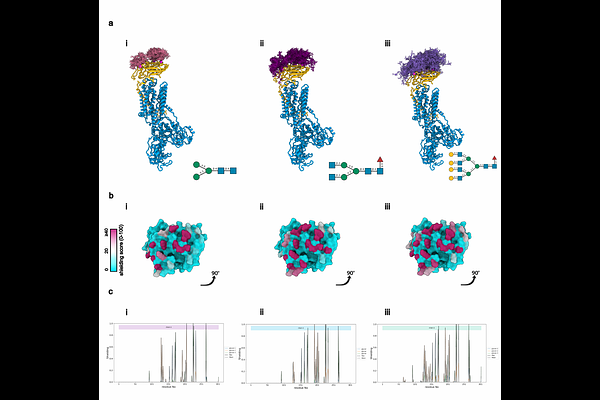
AbstractThe cell surface localization of the Na,K-ATPase (sodium pump) is required for maintaining transmembrane electrochemical gradients. While glycosylation of the {beta}1 subunit facilitates trafficking from the endoplasmic reticulum to the plasma membrane, its role in nanoscale surface organization is not characterized. This study employed GlycoSHIELD computational modeling and DNA-PAINT single-molecule localization microscopy (SMLM) to evaluate how N-glycans influence pump distribution. In-silico simulations indicated that N-glycans sequester the protein core, providing a steric shield that increases with structural complexity. To investigate this experimentally, glycosylation-deficient mutants (3NQ) were generated and confirmed via immunoblotting. Quantitative SMLM analysis of A498 cells demonstrated that wild-type pumps exhibit higher localization density and form larger (144 nm) and more frequent clusters than 3NQ mutants (109 nm). These results indicate that N-glycosylation promotes stable enzyme clustering, supporting a galectin-lattice mechanism of organization rather than steric repulsion.