Symmetric Self-play Online Preference Optimization for Protein Inverse Folding
Symmetric Self-play Online Preference Optimization for Protein Inverse Folding
Zeng, W.; Li, X.; Zou, H.; Dou, Y.; Zhao, X.; Peng, S.
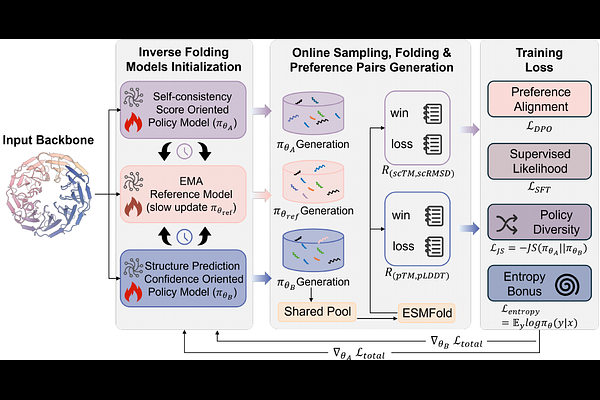
AbstractMulti-objective reinforcement learning based on predicted structure feedback has been introduced into protein inverse folding. However, existing methods typically rely on a single model to optimize multiple structural objectives via a scalarized reward, which can bias the optimization toward dominant objectives and limit the exploration of diverse solutions. Here, we propose a online Symmetric Self-play Preference Optimization (SSP) framework that decouples the optimization of multiple structural objectives by training separate preference models with distinct reward signals, while enabling interaction through a shared sampling pool. This design allows the models to explore diverse optimization trajectories without enforcing a single dominant direction. Extensive experiments on both natural and de novo binder backbone inverse folding tasks demonstrate that SSP consistently improves sequence design self-consistency compared to single-model and existing baselines. Further analysis shows that different structural objectives are only partially aligned and induce distinct optimization directions, as evidenced by metric correlation and white-box analyses. This supports the effectiveness of decoupling objectives to enable higher design quality in protein design.