A deterministic computational kernel encoded in the human genome
A deterministic computational kernel encoded in the human genome
Levy, J.
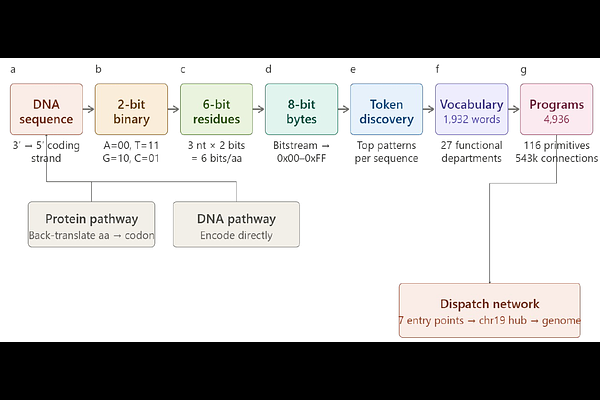
AbstractA computational kernel is defined by four properties: initialization from raw input, a fixed instruction set, a process table with memory organization, and inter-component signal dispatch. Here we show that the human genome satisfies all four. Applying a deterministic 6-bit encoding to the complete human proteome (83,587 isoforms from 32,281 genes) and all chromosome assemblies, we extract 1,932 recurring byte-level vocabulary patterns mapping to 27 functional categories, identify 4,936 genome programs with chromosome-based memory segments, and trace a dispatch network of 543,554 edges routed through a single relay hub. All seven network entry points converge on the mitochondrial genome, which functions as a read-only boot origin. Five null model tests, fifteen robustness analyses, and independent validation against DepMap gene essentiality and STRING protein-protein interactions confirm that this architecture is a property of the genome itself. Vocabulary conservation tracks evolutionary divergence across nine species. The same pipeline applied to composition-matched random sequences fails all four properties.