Mapping spatial cell-cell communication programs by tailoring chains of cells for transformer neural networks
Mapping spatial cell-cell communication programs by tailoring chains of cells for transformer neural networks
Brunn, N.; Guitart, L. C.; Farhadyar, K.; Fullio, C. L.; Kailer, J.; Vogel, T.; Hackenberg, M.; Binder, H.
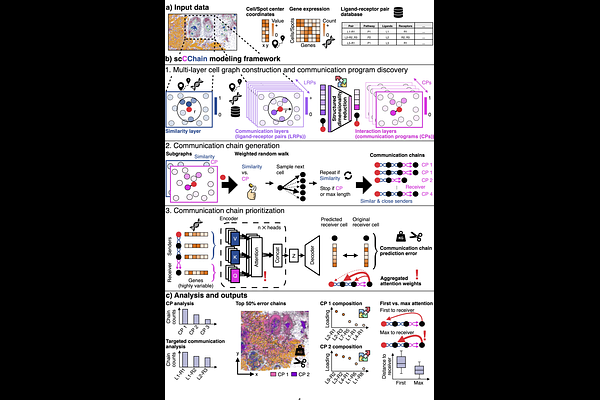
AbstractRecent advances in spatial transcriptomics and computational modeling enable the study of cellular interactions in situ. However, existing methods quantify ligand-receptor activity pairwise or between predefined cell groups, yielding overlapping signals and limited ability to summarize concurrent interactions into programs while localizing communication hotspots. We introduce scCChain, a transformer-based framework that integrates ligand-receptor activity into spatially resolved communication programs and localizes hotspots at spot and single-cell resolution. scCChain first derives candidate programs using structured dimensionality reduction. Subsequently, it samples program-specific communication chains by linking transcriptionally similar sender cells to candidate receivers via weighted random walks on a distance-informed cell graph, borrowing signal from similar neighbors. Transformer-based modeling then scores chains to prioritize communication programs and pinpoint hotspots across the tissue. Applied to human breast cancer spatial transcriptomics data at spot and single-cell resolution, scCChain supports both exploratory communication program discovery and targeted analysis of user-specified ligand-receptor pairs. In spot-level data, it prioritizes a tumor-associated program enriched for pro-angiogenic signaling that localizes to invasive regions. In imaging-based data, it highlights CXCL12-CXCR4 communication hotspots at cellular resolution. Here, we demonstrate that chain-based transformer modeling enables interpretable discovery and mapping of biological meaningful spatial communication programs within complex tissues.