Decoupling Lineage and Intrinsic Information in Single-Cell Lineage Tracing Data with Deep Disentangled Representation Learning
Decoupling Lineage and Intrinsic Information in Single-Cell Lineage Tracing Data with Deep Disentangled Representation Learning
Wen, Y.; Xiong, J.; Gong, F.; Ma, L.; Wan, L.
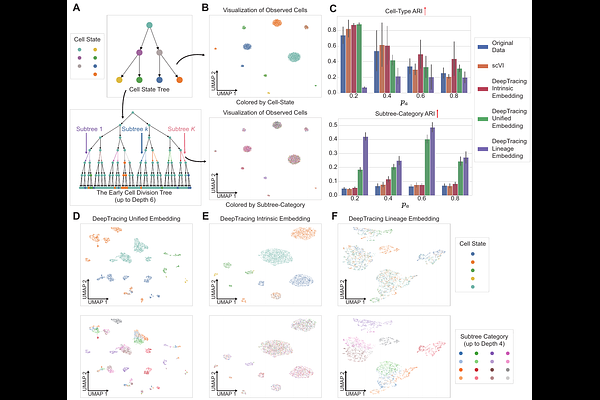
AbstractSingle-cell RNA sequencing combined with lineage tracing technologies provides rich opportunities to study development and tumor evolution, yet existing computational methods struggle to disentangle intrinsic transcriptional states from lineage-driven effects. We introduce DeepTracing, a deep generative framework that integrates disentangled representation learning with lineage-aware Gaussian processes to explicitly separate intrinsic cellular variation from lineage constraints. The model constructs a layered latent space and enforces independence via Total Correlation regularization, producing intrinsic, lineage, and unified embeddings. Across extensive benchmarks, DeepTracing consistently outperforms existing approaches. In TedSim simulations, it achieves superior clustering of cell states and effectively recovers phylogenetic structure, surpassing original expression and scVI. Applied to mouse tumor lineage-tracing data, DeepTracing attains higher ARI/NMI for tumor-type classification than scVI and PORCELAN, accurately separating primary and metastatic tumors and recovering known trajectories such as early lymph-node divergence and liver-to-kidney cross-seeding. In larger datasets, it maintains strong performance while preserving both transcriptomic continuity and lineage fidelity. DeepTracing also reconstructs continuous developmental trajectories in mouse ventral midbrain, isolating temporal effects from intrinsic differentiation. These results establish DeepTracing as a scalable and interpretable framework for analyzing multimodal single-cell data in tumor progression.