BioWorldModel: a single architecture predictsphenotype from genotype across four kingdoms of life
BioWorldModel: a single architecture predictsphenotype from genotype across four kingdoms of life
Shaik, K. H. B.; Sahu, A.
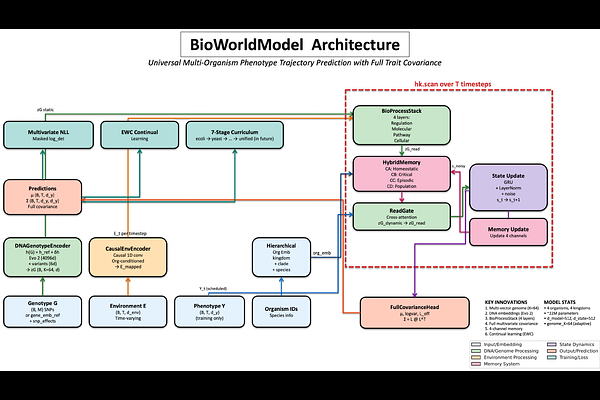
AbstractThe same genome produces different phenotypes in different conditions--yet predictive models encode genotype once and treat each trait independently. Here we show that representing phenotype generation as a dynamic biological process transforms predictive accuracy across bacteria, fungi, animals and plants. BioWorldModel learns how organisms interpret their genome: frozen gene embeddings (species context) modulated by individual variation pass through four biological process layers (regulation to expression to pathway to cellular) that respond to environment and time. A state-conditioned attention mechanism rereads this dynamic representation, predicting full multivariate trait distributions. Without modification, the architecture achieves mean correlation r = 0.678 on 214 bacterial growth traits (207% better than ridge regression), r = 0.915 on 35 yeast fitness traits (167% better), r = 0.499 on 199 fly phenotypes in small-sample regime (760% better), and r = 0.995 on 36 rice traits (49% better). Ablations confirm that modeling biological process--not model size--drives performance. When neural architectures represent how biology generates phenotype rather than merely associating genotype with outcome, they capture what static methods miss.