Benchmarking niche identification via domain segmentation for spatial transcriptomics data
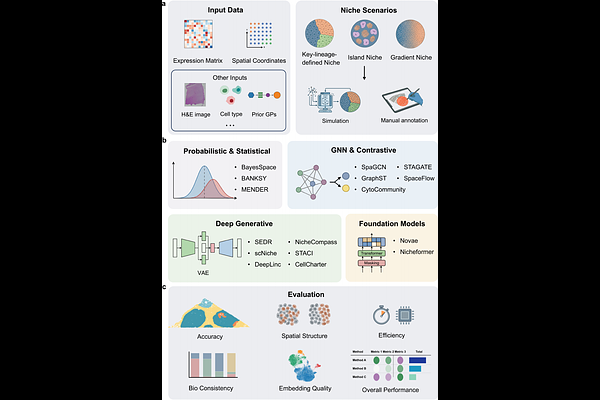
Benchmarking niche identification via domain segmentation for spatial transcriptomics data
Wang, Y.; Chen, Y.; Yang, L.; Wang, C.; Cai, J.; Xin, H.
AbstractTissue niches are spatially organized microenvironments in which coordinated multicellular interactions shape cellular states and biological functions. Currently, niche identification is routinely performed using domain segmentation frameworks. While interrelated, spatial domains and niches are not fundamentally equivalent. The former emphasizes intra-domain compositional consistency and transcriptomic homogeneity, whereas the latter is defined by the emergent properties of localized signaling gradients and the functional reciprocity between key cell lineages. Here, we present a high-resolution reference by thoroughly annotating single-cell resolution CosMx ST data of a human follicular lymphoid hyperplasia lymph node, a dynamic, non-compartmentalized tissue containing several critical immune niches defined by specific lineage architectures. We systematically benchmarked 16 contemporary domain segmentation algorithms, demonstrating that most methods in their default configurations fail to recapitulate biologically defined niche boundaries. Our analysis reveals that the definitive, disjoint spatial distributions of key functional lineages are frequently obscured by the stochastic infiltration of peripheral cell types. Such reduction in the spatial signal-to-noise ratio represents a primary bottleneck for existing algorithms, which prioritize local transcriptomic variance over global architectural logic. Following this observation, we demonstrate that strategic weighting of core functional lineages can restore the resolution of spatial niches in select domain segmentation frameworks. Cross-comparison against compartmentalized tissues further underscores the unique challenges of niche identification in non-mechanically separated environments and clarifies the fundamental divergence between structural domain segmentation and functional niche discovery. Our work delineates the limitations of current paradigms and advocates for the development of specialized computational approaches tailored specifically to the complexity of functional microenvironments.