Dynamic engagement of dual-role regulators by the Sin3 complex
Dynamic engagement of dual-role regulators by the Sin3 complex
Olivet, J.; Shewakramani, N. R.; Cesare, J.; Laval, F.; Nde, J.; Van de Veire, J.; Brammerloo, Y.; Brebel, B.; Debnath, O.; Richardson, A. D.; Yue, H.; Wang, Y.; Spirohn-Fitzgerald, K.; Lemmens, I. H.; Hao, T.; Tavernier, J.; Calderwood, M. A.; Seo, H.-S.; Roth, F. P.; Dhe-Paganon, S.; Twizere, J.-C.; Hill, D. E.; Vidal, M.; Washburn, M. P.; Das, K.
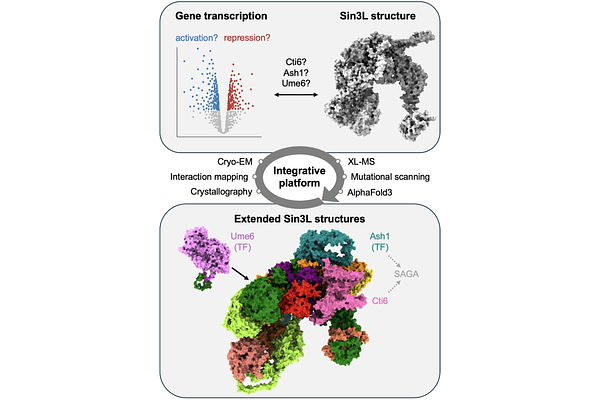
AbstractGene expression is governed by dynamic switches between repressive and activating transcriptional states. Among the molecules mediating these transitions, chromatin readers and transcription factors play pivotal roles. However, how they assemble with regulatory machineries to enable crosstalk between gene repression and activation remains unknown. Here, we use an integrative structural dynamics approach - combining cryo-EM, crosslinking mass spectrometry, fragment-resolved protein interactome mapping and crystallography - to show how the dual-role chromatin reader Cti6 and transcription factors Ash1 and Ume6 engage the Sin3 deacetylase complex, a major regulatory hub in eukaryotes. We find that Cti6 competes with Ash1 to drive its dynamic recruitment to a shared peripheral module, while Ume6 engages the Sin3 scaffold through a defined, minimal interface. Using high-throughput mutational scanning, we reveal deleterious and gain-of-function mutations in Sin3, identifying evolutionarily conserved residues essential for anchoring transcription factors. Together, these results provide structural and functional insights into how dual-role regulators engage the central Sin3 complex, revealing subtle assembly principles that may facilitate crosstalk between gene repression and activation. They also establish an integrative multidisciplinary framework to dissect the dynamics of macromolecular assemblies across biological systems.